 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | GSVIVT01007564001 | ||||||||
| Common Name | VIT_17s0000g09980 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; rosids incertae sedis; Vitales; Vitaceae; Vitis
|
||||||||
| Family | C2H2 | ||||||||
| Protein Properties | Length: 950aa MW: 105704 Da PI: 8.3751 | ||||||||
| Description | C2H2 family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | zf-C2H2 | 12.5 | 0.00045 | 833 | 858 | 1 | 23 |
EEET..TTTEEESSHHHHHHHHHH.T CS
zf-C2H2 1 ykCp..dCgksFsrksnLkrHirt.H 23
y+C C++sFs+k +L H ++ +
GSVIVT01007564001 833 YQCDmeGCTMSFSSKPELALHKKNiC 858
99*******************99866 PP
| |||||||
| 2 | zf-C2H2 | 13.6 | 0.00019 | 858 | 880 | 3 | 23 |
ET..TTTEEESSHHHHHHHHHHT CS
zf-C2H2 3 Cp..dCgksFsrksnLkrHirtH 23
Cp Cgk F ++ +L++H r+H
GSVIVT01007564001 858 CPvkGCGKKFFSHKYLVQHRRVH 880
9999*****************99 PP
| |||||||
| 3 | zf-C2H2 | 11.9 | 0.00069 | 916 | 942 | 1 | 23 |
EEET..TTTEEESSHHHHHHHHHH..T CS
zf-C2H2 1 ykCp..dCgksFsrksnLkrHirt..H 23
y C+ Cg++F+ s++ rH r+ H
GSVIVT01007564001 916 YICTeaGCGQTFRFVSDFSRHKRKtgH 942
899999****************99666 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| SMART | SM00558 | 2.1E-44 | 1 | 162 | IPR003347 | JmjC domain |
| PROSITE profile | PS51184 | 33.633 | 1 | 162 | IPR003347 | JmjC domain |
| SuperFamily | SSF51197 | 1.92E-26 | 8 | 176 | No hit | No description |
| Pfam | PF02373 | 2.0E-37 | 26 | 145 | IPR003347 | JmjC domain |
| SMART | SM00355 | 7.7 | 833 | 855 | IPR015880 | Zinc finger, C2H2-like |
| PROSITE profile | PS50157 | 12.383 | 856 | 885 | IPR007087 | Zinc finger, C2H2 |
| SMART | SM00355 | 0.0045 | 856 | 880 | IPR015880 | Zinc finger, C2H2-like |
| Gene3D | G3DSA:3.30.160.60 | 1.1E-5 | 857 | 879 | IPR013087 | Zinc finger C2H2-type/integrase DNA-binding domain |
| PROSITE pattern | PS00028 | 0 | 858 | 880 | IPR007087 | Zinc finger, C2H2 |
| SuperFamily | SSF57667 | 1.56E-9 | 872 | 914 | No hit | No description |
| Gene3D | G3DSA:3.30.160.60 | 1.0E-9 | 880 | 908 | IPR013087 | Zinc finger C2H2-type/integrase DNA-binding domain |
| PROSITE profile | PS50157 | 10.741 | 886 | 915 | IPR007087 | Zinc finger, C2H2 |
| SMART | SM00355 | 0.0014 | 886 | 910 | IPR015880 | Zinc finger, C2H2-like |
| PROSITE pattern | PS00028 | 0 | 888 | 910 | IPR007087 | Zinc finger, C2H2 |
| SuperFamily | SSF57667 | 7.62E-8 | 904 | 938 | No hit | No description |
| Gene3D | G3DSA:3.30.160.60 | 3.5E-9 | 909 | 939 | IPR013087 | Zinc finger C2H2-type/integrase DNA-binding domain |
| SMART | SM00355 | 0.24 | 916 | 942 | IPR015880 | Zinc finger, C2H2-like |
| PROSITE profile | PS50157 | 11.489 | 916 | 947 | IPR007087 | Zinc finger, C2H2 |
| PROSITE pattern | PS00028 | 0 | 918 | 942 | IPR007087 | Zinc finger, C2H2 |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0009741 | Biological Process | response to brassinosteroid | ||||
| GO:0009826 | Biological Process | unidimensional cell growth | ||||
| GO:0010228 | Biological Process | vegetative to reproductive phase transition of meristem | ||||
| GO:0033169 | Biological Process | histone H3-K9 demethylation | ||||
| GO:0035067 | Biological Process | negative regulation of histone acetylation | ||||
| GO:0048366 | Biological Process | leaf development | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003676 | Molecular Function | nucleic acid binding | ||||
| GO:0046872 | Molecular Function | metal ion binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 950 aa Download sequence Send to blast |
MRGISRAKGS LLRFMKEEIP GVTSPMVYVA MMFSWFAWHV EDHDLHSLNY LHMGAGKTWY 60 GVPREAAVAF EEVVRVHGYG GEINPLVTFA VLGEKTTVMS PEVFVSAGIP CCRLVQNPGE 120 FVVTFPRAYH SGFSHGFNCG EAANIATPEW LRVAKDAAIR RASINYPPMV SHFQLLYDLA 180 LALCSRIPMS ISVEPRSSRL KDKKRGEGET VVKELFVQNI MQNNDLLHIL GKGSSIVLLP 240 KRSSDISVCP NLRVGSSSRV KPRLSLGLCN LEEAMKTSKS ILHLSHGNDN GSALTSQTQN 300 METKIESISH GDGLSDQALF SCVTCGILSF ACVALIQPRE AAARYLMSAD CSFFNDWIVG 360 SGPSGVANED FTGVSGDVHN SELNSCSGWM RKRVPNALFD VPIQSANYQI QTVDQNNEVV 420 SNTGTQKNTS ALGLLALTYA NSSDSEEDQL EPDIPLEADN LASTESNSSE GIFRDPLAIS 480 WATSKYSPVG HDAERAKFSN AIVPVENTNM SFAPRSDEDY SRIHVFCLEH AVEVEQQLRP 540 IGGVNMLLLC HPDYPKVEAE AKLVAEDLGI DYLWNDFVYR DATKEDGEMI QSALDSEECI 600 PGNGDWAVKL GVNLYYSANL SRSPLYIKQM PYNSVIYNVF GRSSANSPTA PDVYGRGPGK 660 QKKIVVAGKW CGKVWMSNQV HPLLAQKDPE EQEEDRNFHV WVKKPDEKPE RKSESSRKAE 720 TSSAPRKSGR KRKMMVENGS TKKANRPERE DPTPRRRNSC EQSAREFDSY VEDELEGGPS 780 TRLRRRNPKP PKELEAKPVV KKQTARKKAK KAPAAKAPGN HNNAKIQDEE EEYQCDMEGC 840 TMSFSSKPEL ALHKKNICPV KGCGKKFFSH KYLVQHRRVH IDDRPLKCPW KGCKMTFKWA 900 WARTEHIRVH TGARPYICTE AGCGQTFRFV SDFSRHKRKT GHSAKKARG* |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 6a57_A | 9e-68 | 833 | 947 | 23 | 137 | Lysine-specific demethylase REF6 |
| 6a58_A | 9e-68 | 833 | 947 | 23 | 137 | Lysine-specific demethylase REF6 |
| 6a59_A | 9e-68 | 833 | 947 | 23 | 137 | Lysine-specific demethylase REF6 |
| Search in ModeBase | ||||||
| Expression -- UniGene ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| UniGene ID | E-value | Expressed in | ||||
| Vvi.29525 | 0.0 | inflorescence | ||||
| Expression -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| Uniprot | TISSUE SPECIFICITY: Highly expressed in the shoot apical meristem and primary and secondary root tips, and lower expression in cotyledons, leaves and root axis along vascular tissues. Detected in inflorescences, stems and siliques. {ECO:0000269|PubMed:15377760, ECO:0000269|PubMed:18713399}. | |||||
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Histone demethylase that demethylates 'Lys-27' (H3K27me) of histone H3. Demethylates both tri- (H3K27me3) and di-methylated (H3K27me2) H3K27me (PubMed:21642989, PubMed:27111035). Demethylates also H3K4me3/2 and H3K36me3/2 in an in vitro assay (PubMed:20711170). Involved in the transcriptional regulation of hundreds of genes regulating developmental patterning and responses to various stimuli (PubMed:18467490). Binds DNA via its four zinc fingers in a sequence-specific manner, 5'-CTCTGYTY-3', to promote the demethylation of H3K27me3 and the regulation of organ boundary formation (PubMed:27111034, PubMed:27111035). Involved in the regulation of flowering time by repressing FLOWERING LOCUS C (FLC) expression (PubMed:15377760). Interacts with the NF-Y complexe to regulate SOC1 (PubMed:25105952). Mediates the recruitment of BRM to its target loci (PubMed:27111034). {ECO:0000269|PubMed:15377760, ECO:0000269|PubMed:18467490, ECO:0000269|PubMed:20711170, ECO:0000269|PubMed:21642989, ECO:0000269|PubMed:25105952, ECO:0000269|PubMed:27111034, ECO:0000269|PubMed:27111035}. | |||||
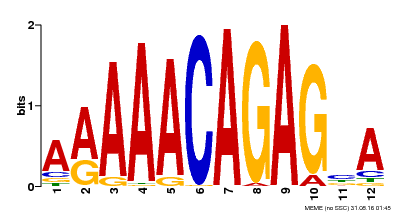
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00608 | ChIP-seq | Transfer from AT3G48430 | Download |
| |||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | AM466832 | 0.0 | AM466832.1 Vitis vinifera, whole genome shotgun sequence, contig VV78X011128.17, clone ENTAV 115. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_028116352.1 | 0.0 | LOW QUALITY PROTEIN: lysine-specific demethylase REF6-like | ||||
| Swissprot | Q9STM3 | 0.0 | REF6_ARATH; Lysine-specific demethylase REF6 | ||||
| TrEMBL | F6GTM2 | 0.0 | F6GTM2_VITVI; Uncharacterized protein | ||||
| STRING | VIT_17s0000g09980.t01 | 0.0 | (Vitis vinifera) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Representative plant | OGRP4401 | 12 | 17 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT3G48430.1 | 0.0 | relative of early flowering 6 | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | GSVIVT01007564001 |



