 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | GSMUA_Achr8P07460_001 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; Liliopsida; Petrosaviidae; commelinids; Zingiberales; Musaceae; Musa
|
||||||||
| Family | C2H2 | ||||||||
| Protein Properties | Length: 301aa MW: 35187.1 Da PI: 8.9081 | ||||||||
| Description | C2H2 family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | zf-C2H2 | 20.7 | 1.1e-06 | 28 | 52 | 1 | 23 |
EEET..TTTEEESSHHHHHHHHHHT CS
zf-C2H2 1 ykCp..dCgksFsrksnLkrHirtH 23
++Cp dC s++rk++L+rH+ tH
GSMUA_Achr8P07460_001 28 FSCPvdDCRVSYRRKDHLTRHLLTH 52
89*********************99 PP
| |||||||
| 2 | zf-C2H2 | 15.5 | 4.9e-05 | 57 | 82 | 1 | 23 |
EEET..TTTEEESSHHHHHHHHHH.T CS
zf-C2H2 1 ykCp..dCgksFsrksnLkrHirt.H 23
+ Cp +C++ F k n+krH+r H
GSMUA_Achr8P07460_001 57 FACPvdNCNRKFGIKANMKRHMREiH 82
78*********************988 PP
| |||||||
| 3 | zf-C2H2 | 20.7 | 1.1e-06 | 94 | 118 | 1 | 23 |
EEET..TTTEEESSHHHHHHHHHHT CS
zf-C2H2 1 ykCp..dCgksFsrksnLkrHirtH 23
y+C+ Cg++F+ +s Lk+H H
GSMUA_Achr8P07460_001 94 YVCGesGCGRTFKYPSKLKKHEEAH 118
99*******************8776 PP
| |||||||
| 4 | zf-C2H2 | 14.3 | 0.00012 | 129 | 151 | 3 | 23 |
ET.TTTEEESSHHHHHHHHHH.T CS
zf-C2H2 3 Cp.dCgksFsrksnLkrHirt.H 23
C+ C+k+F++ + Lk H rt H
GSMUA_Achr8P07460_001 129 CEpGCTKTFTNAECLKAHVRTcH 151
668******************66 PP
| |||||||
| 5 | zf-C2H2 | 20.5 | 1.3e-06 | 185 | 209 | 2 | 23 |
EET..TTTEEESSHHHHHHHHHH.T CS
zf-C2H2 2 kCp..dCgksFsrksnLkrHirt.H 23
+C+ dC+ +Fs+ snL++Hir H
GSMUA_Achr8P07460_001 185 ECNfkDCNCTFSNASNLRQHIRAiH 209
69999*****************888 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| SuperFamily | SSF57667 | 5.04E-8 | 21 | 54 | No hit | No description |
| Gene3D | G3DSA:3.30.160.60 | 4.5E-10 | 24 | 54 | IPR013087 | Zinc finger C2H2-type/integrase DNA-binding domain |
| SMART | SM00355 | 0.0045 | 28 | 52 | IPR015880 | Zinc finger, C2H2-like |
| PROSITE profile | PS50157 | 9.91 | 28 | 57 | IPR007087 | Zinc finger, C2H2 |
| PROSITE pattern | PS00028 | 0 | 30 | 52 | IPR007087 | Zinc finger, C2H2 |
| SuperFamily | SSF57667 | 1.81E-6 | 55 | 83 | No hit | No description |
| Gene3D | G3DSA:3.30.160.60 | 6.6E-7 | 55 | 84 | IPR013087 | Zinc finger C2H2-type/integrase DNA-binding domain |
| SMART | SM00355 | 9.2E-4 | 57 | 82 | IPR015880 | Zinc finger, C2H2-like |
| PROSITE profile | PS50157 | 13.069 | 57 | 87 | IPR007087 | Zinc finger, C2H2 |
| PROSITE pattern | PS00028 | 0 | 59 | 82 | IPR007087 | Zinc finger, C2H2 |
| SuperFamily | SSF57667 | 1.04E-5 | 90 | 119 | No hit | No description |
| Gene3D | G3DSA:3.30.160.60 | 1.0E-7 | 90 | 118 | IPR013087 | Zinc finger C2H2-type/integrase DNA-binding domain |
| SMART | SM00355 | 0.009 | 94 | 118 | IPR015880 | Zinc finger, C2H2-like |
| PROSITE profile | PS50157 | 10.783 | 94 | 118 | IPR007087 | Zinc finger, C2H2 |
| PROSITE pattern | PS00028 | 0 | 96 | 118 | IPR007087 | Zinc finger, C2H2 |
| SMART | SM00355 | 0.18 | 126 | 151 | IPR015880 | Zinc finger, C2H2-like |
| PROSITE profile | PS50157 | 8.6 | 127 | 151 | IPR007087 | Zinc finger, C2H2 |
| Gene3D | G3DSA:3.30.160.60 | 7.0E-5 | 127 | 149 | IPR013087 | Zinc finger C2H2-type/integrase DNA-binding domain |
| PROSITE pattern | PS00028 | 0 | 128 | 151 | IPR007087 | Zinc finger, C2H2 |
| SMART | SM00355 | 49 | 154 | 175 | IPR015880 | Zinc finger, C2H2-like |
| SuperFamily | SSF57667 | 3.97E-7 | 166 | 226 | No hit | No description |
| PROSITE profile | PS50157 | 12.321 | 184 | 214 | IPR007087 | Zinc finger, C2H2 |
| SMART | SM00355 | 0.0021 | 184 | 209 | IPR015880 | Zinc finger, C2H2-like |
| Gene3D | G3DSA:3.30.160.60 | 3.6E-5 | 185 | 211 | IPR013087 | Zinc finger C2H2-type/integrase DNA-binding domain |
| PROSITE pattern | PS00028 | 0 | 186 | 209 | IPR007087 | Zinc finger, C2H2 |
| Gene3D | G3DSA:3.30.160.60 | 2.0E-4 | 212 | 237 | IPR013087 | Zinc finger C2H2-type/integrase DNA-binding domain |
| SMART | SM00355 | 2.8 | 215 | 241 | IPR015880 | Zinc finger, C2H2-like |
| PROSITE profile | PS50157 | 9.162 | 215 | 241 | IPR007087 | Zinc finger, C2H2 |
| PROSITE pattern | PS00028 | 0 | 217 | 241 | IPR007087 | Zinc finger, C2H2 |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0005730 | Cellular Component | nucleolus | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| GO:0008097 | Molecular Function | 5S rRNA binding | ||||
| GO:0046872 | Molecular Function | metal ion binding | ||||
| GO:0080084 | Molecular Function | 5S rDNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 301 aa Download sequence Send to blast |
MDQYQEGGTS CEAPSRKPQS RNFNERPFSC PVDDCRVSYR RKDHLTRHLL THQGKLFACP 60 VDNCNRKFGI KANMKRHMRE IHEDESPCEG QKQYVCGESG CGRTFKYPSK LKKHEEAHVK 120 MDCSEVVCCE PGCTKTFTNA ECLKAHVRTC HQYAECEVCG TKQLKRNYKR HELKHELNEK 180 TDRIECNFKD CNCTFSNASN LRQHIRAIHE KMRPFTCQFS GCGKKFPYKH VRDKHEKSCV 240 HDHVQGDFIE TDERLRSRPR GGRKRKCMSV ETLQRKRVVP PSRASVLDDG VNYLRWLLNE 300 Q |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 1tf6_A | 2e-12 | 24 | 152 | 39 | 189 | PROTEIN (TRANSCRIPTION FACTOR IIIA) |
| 1tf6_D | 2e-12 | 24 | 152 | 39 | 189 | PROTEIN (TRANSCRIPTION FACTOR IIIA) |
| Search in ModeBase | ||||||
| Nucleic Localization Signal ? help Back to Top | |||
|---|---|---|---|
| No. | Start | End | Sequence |
| 1 | 257 | 265 | RPRGGRKRK |
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Essential protein (PubMed:22353599). Isoform 1 is a transcription activator the binds both 5S rDNA and 5S rRNA and stimulates the transcription of 5S rRNA gene (PubMed:12711688, PubMed:22353599). Isoform 1 regulates 5S rRNA levels during development (PubMed:22353599). {ECO:0000269|PubMed:12711688, ECO:0000269|PubMed:22353599}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00229 | DAP | Transfer from AT1G72050 | Download |
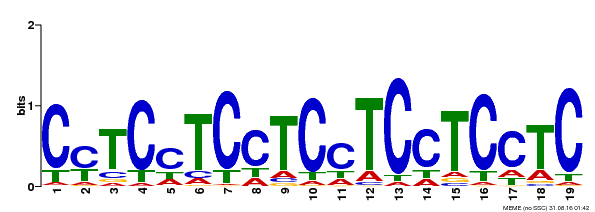
| |||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_009411993.1 | 0.0 | PREDICTED: transcription factor IIIA | ||||
| Swissprot | Q84MZ4 | 5e-97 | TF3A_ARATH; Transcription factor IIIA | ||||
| TrEMBL | M0TNE9 | 0.0 | M0TNE9_MUSAM; Uncharacterized protein | ||||
| STRING | GSMUA_Achr8P07460_001 | 0.0 | (Musa acuminata) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Monocots | OGMP2382 | 38 | 82 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT1G72050.1 | 2e-91 | transcription factor IIIA | ||||



