 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | GSMUA_Achr5P06550_001 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; Liliopsida; Petrosaviidae; commelinids; Zingiberales; Musaceae; Musa
|
||||||||
| Family | HD-ZIP | ||||||||
| Protein Properties | Length: 839aa MW: 92720.5 Da PI: 6.5744 | ||||||||
| Description | HD-ZIP family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | Homeobox | 59.9 | 4.1e-19 | 19 | 77 | 3 | 57 |
--SS--HHHHHHHHHHHHHSSS--HHHHHHHHHHC....TS-HHHHHHHHHHHHHHHHC CS
Homeobox 3 kRttftkeqleeLeelFeknrypsaeereeLAkkl....gLterqVkvWFqNrRakekk 57
k ++t+eq+e+Le+l++++++ps +r++L +++ +++ +q+kvWFqNrR +ek+
GSMUA_Achr5P06550_001 19 KYVRYTPEQVEALERLYHECPKPSSMRRQQLIRECpilsNIEPKQIKVWFQNRRCREKQ 77
5679*****************************************************97 PP
| |||||||
| 2 | START | 164.5 | 7.8e-52 | 163 | 371 | 2 | 205 |
HHHHHHHHHHHHHHHC-TT-EEEEEXCCTTEEEEEEESSS.SCEEEEEEEECCSCHHHHHHHHHCCCGGCT-TT-SEEEEEEEECTT. CS
START 2 laeeaaqelvkkalaeepgWvkssesengdevlqkfeeskvdsgealrasgvvdmvlallveellddkeqWdetlakaetlevissg. 88
+aee+++e+++ka+ ++ Wv+++ +++g++++ +++ s++++g a+ra+g+v +++ v+e+l+d++ W ++++++e+++v+ +g
GSMUA_Achr5P06550_001 163 IAEETLTEFLSKATGTAIEWVQMPGMKPGPDSIGIVAISHGCTGVAARACGLVGLEPT-KVAEILKDRLSWFRDCRSVEIINVLPTGs 249
789*******************************************************.8999999999******************* PP
.EEEEEEEEXXTTXX-SSX.EEEEEEEEEEE.TTS-EEEEEEEEE-TTS--....-TTSEE-EESSEEEEEEEECTCEEEEEEEE-EE CS
START 89 .galqlmvaelqalsplvp.RdfvfvRyirqlgagdwvivdvSvdseqkppe...sssvvRaellpSgiliepksnghskvtwvehvd 171
g+++l +++ +a+++l+p Rdf+ +Ry+ l++ ++v++++S++s+ p+ +++vRae+lpSg+li+p+++g+s +++v+h d
GSMUA_Achr5P06550_001 250 nGTIELLYMQVYAPTTLAPaRDFWLLRYTSILEDRSLVVCERSLNSTLGGPSmptVQPFVRAEMLPSGYLIRPCEGGGSVIHIVDHLD 337
**********************************************99999888789******************************* PP
--SSXXHHHHHHHHHHHHHHHHHHHHHHTXXXXX CS
START 172 lkgrlphwllrslvksglaegaktwvatlqrqce 205
l+ ++++++lr+l++s+++ +kt++ +l+++++
GSMUA_Achr5P06550_001 338 LEPWSVPEVLRPLYESSTVLSQKTTMVALRHLRQ 371
**************************99998875 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| SMART | SM00389 | 1.3E-16 | 16 | 82 | IPR001356 | Homeobox domain |
| CDD | cd00086 | 4.02E-17 | 17 | 79 | No hit | No description |
| Pfam | PF00046 | 9.0E-17 | 19 | 77 | IPR001356 | Homeobox domain |
| SuperFamily | SSF46689 | 4.71E-17 | 19 | 82 | IPR009057 | Homeodomain-like |
| Gene3D | G3DSA:1.10.10.60 | 9.9E-19 | 21 | 77 | IPR009057 | Homeodomain-like |
| PROSITE profile | PS50071 | 15.451 | 22 | 78 | IPR001356 | Homeobox domain |
| CDD | cd14686 | 7.60E-7 | 71 | 110 | No hit | No description |
| PROSITE profile | PS50848 | 24.897 | 153 | 354 | IPR002913 | START domain |
| CDD | cd08875 | 5.78E-76 | 157 | 373 | No hit | No description |
| Gene3D | G3DSA:3.30.530.20 | 7.1E-22 | 160 | 368 | IPR023393 | START-like domain |
| SMART | SM00234 | 1.9E-34 | 162 | 372 | IPR002913 | START domain |
| SuperFamily | SSF55961 | 5.22E-36 | 162 | 372 | No hit | No description |
| Pfam | PF01852 | 1.6E-49 | 163 | 371 | IPR002913 | START domain |
| Pfam | PF08670 | 1.1E-49 | 696 | 838 | IPR013978 | MEKHLA |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0009965 | Biological Process | leaf morphogenesis | ||||
| GO:0010014 | Biological Process | meristem initiation | ||||
| GO:0010075 | Biological Process | regulation of meristem growth | ||||
| GO:0010087 | Biological Process | phloem or xylem histogenesis | ||||
| GO:0048263 | Biological Process | determination of dorsal identity | ||||
| GO:0080060 | Biological Process | integument development | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| GO:0008289 | Molecular Function | lipid binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 839 aa Download sequence Send to blast |
MIALTSCEEG KMGIVDPGKY VRYTPEQVEA LERLYHECPK PSSMRRQQLI RECPILSNIE 60 PKQIKVWFQN RRCREKQRKE AARLQAVNRK LTAMNKLLME ENDRLQKQVS QMVYENGYFR 120 QQTQNAALTT TNASCESVVT NGQQHLTPQH PPRDASPAGL MSIAEETLTE FLSKATGTAI 180 EWVQMPGMKP GPDSIGIVAI SHGCTGVAAR ACGLVGLEPT KVAEILKDRL SWFRDCRSVE 240 IINVLPTGSN GTIELLYMQV YAPTTLAPAR DFWLLRYTSI LEDRSLVVCE RSLNSTLGGP 300 SMPTVQPFVR AEMLPSGYLI RPCEGGGSVI HIVDHLDLEP WSVPEVLRPL YESSTVLSQK 360 TTMVALRHLR QIAHETTHSS VSGWGKRPAA LRALSQRLSR GFNEAVNGFV DDGWSMVSSD 420 GMEDITVLVN SSCSKLMGLN LSFVNGCPST TSSVLCAKAT MLLQNISPPM LLRFLREHRS 480 EWADSNIDGY SAASVKAIPH TLPISRTGCS GGQVILPLAH TLEHEEFLEV IKFESIGNNR 540 EMPMPRDLYL LQLCNGVDEN AVGTCSELIF APIDASFADD APLLPSGFRI IPLDFKMDTN 600 SPNRTLDLAS ALEVGPTGSR VHNDYSGNRG NIRSVMTIAF QFAFESHLQE DVASMARQYI 660 RNIIASVQRL SLTLSPSHLG SHGGLRIPPG SPEAATLACW ICHSYRSYSG AEFLKPSADN 720 SDSLLKMLWH HSDAIICCSL KAMPVFTFAN QAGLDMLETT LIALQDITLE KIFVDEGRKM 780 ICAELPHVIQ QGSICLQAGL CISSMGRPVS YERAVAWKVL DNEDNAHCIC FMFVNWSFV |
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Probable transcription factor involved in the regulation of meristem development to promote lateral organ formation. May regulates procambial and vascular tissue formation or maintenance, and vascular development in inflorescence stems. {ECO:0000269|PubMed:15598805, ECO:0000269|PubMed:15705957, ECO:0000269|PubMed:16617092}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00200 | DAP | Transfer from AT1G52150 | Download |
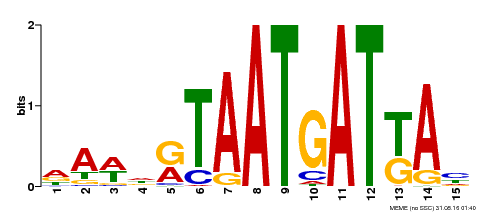
| |||
| Regulation -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | INDUCTION: By auxin. Repressed by miR165 and miR166. {ECO:0000269|PubMed:15773855, ECO:0000269|PubMed:16033795, ECO:0000269|PubMed:16617092, ECO:0000269|PubMed:17237362}. | |||||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_009399812.1 | 0.0 | PREDICTED: homeobox-leucine zipper protein ATHB-15-like isoform X1 | ||||
| Swissprot | Q9ZU11 | 0.0 | ATB15_ARATH; Homeobox-leucine zipper protein ATHB-15 | ||||
| TrEMBL | M0SW84 | 0.0 | M0SW84_MUSAM; Uncharacterized protein | ||||
| STRING | GSMUA_Achr5P06550_001 | 0.0 | (Musa acuminata) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Monocots | OGMP374 | 38 | 197 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT1G52150.1 | 0.0 | HD-ZIP family protein | ||||
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



