 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | GSMUA_Achr4P23160_001 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; Liliopsida; Petrosaviidae; commelinids; Zingiberales; Musaceae; Musa
|
||||||||
| Family | G2-like | ||||||||
| Protein Properties | Length: 272aa MW: 29937.8 Da PI: 9.3261 | ||||||||
| Description | G2-like family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | G2-like | 100.9 | 8.1e-32 | 69 | 122 | 2 | 55 |
G2-like 2 prlrWtpeLHerFveaveqLGGsekAtPktilelmkvkgLtlehvkSHLQkYRl 55
pr+rWt++LH++Fv+av+ LGG+e+AtPk++lelm+vk+Ltl+hvkSHLQ+YR+
GSMUA_Achr4P23160_001 69 PRMRWTSTLHAHFVHAVKLLGGHERATPKSVLELMNVKDLTLAHVKSHLQMYRT 122
9****************************************************7 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| SuperFamily | SSF46689 | 1.56E-15 | 66 | 123 | IPR009057 | Homeodomain-like |
| Gene3D | G3DSA:1.10.10.60 | 3.5E-28 | 67 | 123 | IPR009057 | Homeodomain-like |
| TIGRFAMs | TIGR01557 | 1.9E-23 | 69 | 122 | IPR006447 | Myb domain, plants |
| Pfam | PF00249 | 4.7E-7 | 70 | 121 | IPR001005 | SANT/Myb domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0006355 | Biological Process | regulation of transcription, DNA-templated | ||||
| GO:0080060 | Biological Process | integument development | ||||
| GO:0005618 | Cellular Component | cell wall | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0044212 | Molecular Function | transcription regulatory region DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 272 aa Download sequence Send to blast |
MSTIYPDLSL HISPPSVSLN HAEPRLSLGL QSPGSSLSNH HEKLHRPQIC GFKRSSRSAH 60 GGKRSVRAPR MRWTSTLHAH FVHAVKLLGG HERATPKSVL ELMNVKDLTL AHVKSHLQMY 120 RTVKSTDRGE AGQGQADMGI SQRTGMEEVE GGLPCDKAGS EITSSYHSLS TPTPPTTQVK 180 SPREMYPSAE RHAWNPSIKQ NGLAYPFFRS DDLLSNDYQA LVEDQPQALT QSQEQRLNLV 240 PNSIGAAHTK MPDLEISLGS HDASKMLVNY IA |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 6j4k_A | 6e-17 | 70 | 124 | 4 | 58 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j4k_B | 6e-17 | 70 | 124 | 4 | 58 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j5b_A | 6e-17 | 70 | 124 | 4 | 58 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j5b_C | 6e-17 | 70 | 124 | 4 | 58 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j5b_D | 6e-17 | 70 | 124 | 4 | 58 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j5b_F | 6e-17 | 70 | 124 | 4 | 58 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j5b_H | 6e-17 | 70 | 124 | 4 | 58 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j5b_J | 6e-17 | 70 | 124 | 4 | 58 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| Search in ModeBase | ||||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00107 | PBM | Transfer from AT5G42630 | Download |
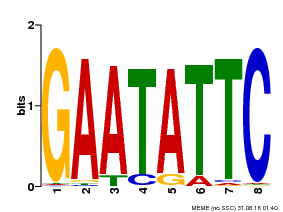
| |||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | - | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | AC226043 | 1e-114 | AC226043.1 Musa acuminata clone BAC MA4-57L19, complete sequence. | |||
| GenBank | AC226044 | 1e-114 | AC226044.1 Musa acuminata clone BAC MA4-69C10, complete sequence. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_009398300.1 | 0.0 | PREDICTED: probable transcription factor KAN4 isoform X1 | ||||
| TrEMBL | M0SRC7 | 0.0 | M0SRC7_MUSAM; Uncharacterized protein | ||||
| STRING | GSMUA_Achr4P23160_001 | 0.0 | (Musa acuminata) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Monocots | OGMP3742 | 31 | 68 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT5G42630.2 | 2e-49 | G2-like family protein | ||||



