 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | GSBRNA2T00149707001 | ||||||||
| Common Name | GSBRNA2T00149707001 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; malvids; Brassicales; Brassicaceae; Brassiceae; Brassica
|
||||||||
| Family | CAMTA | ||||||||
| Protein Properties | Length: 1029aa MW: 114916 Da PI: 5.3884 | ||||||||
| Description | CAMTA family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | CG-1 | 180.6 | 1.8e-56 | 20 | 136 | 3 | 118 |
CG-1 3 ke.kkrwlkneeiaaiLenfekheltlelktrpksgsliLynrkkvryfrkDGyswkkkkdgktvrEdhekLKvggvevlycyYahseen 91
+e ++rwl++ ei++iL+n++k+++++e++t+p sgs++L++rk++ryfrkDG++w+kk+dgktv+E+he+LK g+v+vl+cyYah+++n
GSBRNA2T00149707001 20 SEaRNRWLRPPEICEILQNYQKFQISTEPPTTPASGSVFLFDRKVLRYFRKDGHNWRKKRDGKTVKEAHERLKAGSVDVLHCYYAHGQDN 109
4449************************************************************************************** PP
CG-1 92 ptfqrrcywlLeeelekivlvhylevk 118
++fqrr+yw+L+eel++iv+vhylevk
GSBRNA2T00149707001 110 EHFQRRSYWMLQEELSHIVFVHYLEVK 136
************************985 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| PROSITE profile | PS51437 | 82.089 | 15 | 141 | IPR005559 | CG-1 DNA-binding domain |
| SMART | SM01076 | 1.1E-82 | 18 | 136 | IPR005559 | CG-1 DNA-binding domain |
| Pfam | PF03859 | 1.9E-49 | 21 | 135 | IPR005559 | CG-1 DNA-binding domain |
| SuperFamily | SSF81296 | 3.5E-16 | 453 | 536 | IPR014756 | Immunoglobulin E-set |
| CDD | cd00204 | 4.04E-11 | 615 | 745 | No hit | No description |
| Gene3D | G3DSA:1.25.40.20 | 2.4E-15 | 616 | 749 | IPR020683 | Ankyrin repeat-containing domain |
| PROSITE profile | PS50297 | 17.184 | 619 | 745 | IPR020683 | Ankyrin repeat-containing domain |
| SuperFamily | SSF48403 | 1.71E-15 | 631 | 748 | IPR020683 | Ankyrin repeat-containing domain |
| PROSITE profile | PS50088 | 9.11 | 686 | 718 | IPR002110 | Ankyrin repeat |
| SuperFamily | SSF52540 | 1.46E-8 | 846 | 899 | IPR027417 | P-loop containing nucleoside triphosphate hydrolase |
| SMART | SM00015 | 0.92 | 848 | 870 | IPR000048 | IQ motif, EF-hand binding site |
| PROSITE profile | PS50096 | 9.029 | 850 | 878 | IPR000048 | IQ motif, EF-hand binding site |
| Pfam | PF00612 | 0.0041 | 851 | 869 | IPR000048 | IQ motif, EF-hand binding site |
| SMART | SM00015 | 0.0011 | 871 | 893 | IPR000048 | IQ motif, EF-hand binding site |
| PROSITE profile | PS50096 | 9.468 | 872 | 896 | IPR000048 | IQ motif, EF-hand binding site |
| Pfam | PF00612 | 1.2E-4 | 874 | 893 | IPR000048 | IQ motif, EF-hand binding site |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0009409 | Biological Process | response to cold | ||||
| GO:0010150 | Biological Process | leaf senescence | ||||
| GO:0042742 | Biological Process | defense response to bacterium | ||||
| GO:0050832 | Biological Process | defense response to fungus | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| GO:0005516 | Molecular Function | calmodulin binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 1029 aa Download sequence Send to blast |
MAEARRFSLN NELDVGQILS EARNRWLRPP EICEILQNYQ KFQISTEPPT TPASGSVFLF 60 DRKVLRYFRK DGHNWRKKRD GKTVKEAHER LKAGSVDVLH CYYAHGQDNE HFQRRSYWML 120 QEELSHIVFV HYLEVKGSRV STSYNRMQRT EDSTQETGEV YTSERNGYAS GSINQYDHSN 180 NQSQATDSAS VNGVHTPELE DAQSAYNQQG SPILYSHQAL QQPPATGFDP YYQMSLTPRD 240 SYQKEIHTIP SSTMVEKSRT INGPVVTNSI KNKKSIDSQT WEEILGNCGS GGEGLPMQPH 300 SEHEGLDQML QSYSFTMQDF ASLQESIVKS QNQELSSGLT SDRSMWFQGQ AVDIEPNALS 360 NLASSEKAPY LSTMKQHLLD GALGEEGLKK MDSFNRWMSK ELGELGDVGV TADANESFTH 420 SSSTAYWEEV ESEDVSNGGY VMSPSLSKEQ LFTIIDFAPN WTYVGCEVKV LVSGKFLKTA 480 ESGEWSCMFG QTEVPADIIA NGILECVAPM HEAGRVPFYV TCSNRLACSE VREFEYKVLE 540 SQGFDRDTDD SSTACNSIES LEARFVKLLC SKSDCTHSSL PGGNDSYLSQ VSEKISLLLF 600 ENDDQLDQML MNEISQENMK NNLLQEALKE SLHSWLLQKI AEGGKGPNVL DEGGQGVLHF 660 AAALGYNWAL EPTIVAGVSV DFRDVNGWTA LHWAAFFGRE LIIGSLIALG ASPGTLTDPN 720 PDFPSGSTPS DLAYANGYKG IAGYLSEYAL RAHVSLLSLN EKNAETSLGA AVEAAPSPSS 780 SALTDSLTAV RNASQAAARI HQVFRAQSFQ KKQMKEFGDR KLGMSEERAL SMLAPKTHRP 840 GRGHSDDSVQ AAAIRIQNKF RGYKGRKDYL ITRQRIIKIQ AHVRGYQVRK NYRKIIWSVG 900 ILEKVILRWR RKGAGLRGFK SDALVTKMQD GTEKEEDDDF FKQGRKQTEE RLEKALARVK 960 SMVQYPEARD QYRRLLNVVN DIQESKVEKA LANSEEATCF DDDLIDIEAL LGDDDTLMMP 1020 MSSTLWNA* |
| Expression -- UniGene ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| UniGene ID | E-value | Expressed in | ||||
| Bna.6884 | 0.0 | flower| leaf| seed | ||||
| Expression -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| Uniprot | TISSUE SPECIFICITY: Expressed in roots, stems, leaves, carpels, and siliques, but not in stigmas or other parts of the flower. {ECO:0000269|PubMed:11162426, ECO:0000269|PubMed:12218065, ECO:0000269|PubMed:14581622}. | |||||
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Transcription activator that binds to the DNA consensus sequence 5'-[ACG]CGCG[GTC]-3'. Binds calmodulin in a calcium-dependent manner in vitro (PubMed:12218065). Regulates transcriptional activity in response to calcium signals (Probable). Involved in freezing tolerance in association with CAMTA1 and CAMTA2 (PubMed:23581962). Required for the cold-induced expression of DREB1B/CBF1, DREB1C/CBF2, ZAT12 and GOLS3 (PubMed:19270186). Involved in response to cold. Contributes together with CAMTA5 to the positive regulation of the cold-induced expression of DREB1A/CBF3, DREB1B/CBF1 and DREB1C/CBF2 (PubMed:28351986). Involved together with CAMTA2 and CAMTA4 in the positive regulation of a general stress response (GSR) (PubMed:25039701). Involved in the regulation of GSR amplitude downstream of MEKK1 (PubMed:25157030). Involved in the regulation of a set of genes involved in defense responses against pathogens (PubMed:18298954). Involved in the regulation of both basal resistance and systemic acquired resistance (SAR) (PubMed:21900483). Acts as negative regulator of plant immunity (PubMed:19122675, PubMed:21900483, PubMed:22345509, PubMed:28407487). Binds to the promoter of the defense-related gene EDS1 and represses its expression (PubMed:19122675). Binds to the promoter of the defense-related gene NDR1 and represses its expression (PubMed:22345509). Involved in defense against insects (PubMed:23072934, PubMed:22371088). Required for tolerance to the generalist herbivore Trichoplusia ni, and contributes to the positive regulation of genes associated with glucosinolate metabolism (PubMed:23072934). Required for tolerance to Bradysia impatiens larvae. Mediates herbivore-induced wound response (PubMed:22371088). Required for wound-induced jasmonate accumulation (PubMed:23072934, PubMed:22371088). Involved in the regulation of ethylene-induced senescence by binding to the promoter of the senescence-inducer gene EIN3 and repressing its expression (PubMed:22345509). {ECO:0000269|PubMed:12218065, ECO:0000269|PubMed:18298954, ECO:0000269|PubMed:19122675, ECO:0000269|PubMed:19270186, ECO:0000269|PubMed:21900483, ECO:0000269|PubMed:22345509, ECO:0000269|PubMed:22371088, ECO:0000269|PubMed:23072934, ECO:0000269|PubMed:23581962, ECO:0000269|PubMed:25039701, ECO:0000269|PubMed:25157030, ECO:0000269|PubMed:28351986, ECO:0000269|PubMed:28407487, ECO:0000305|PubMed:11925432}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00042 | PBM | Transfer from AT2G22300 | Download |
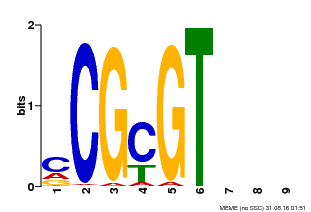
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | GSBRNA2T00149707001 |
| Regulation -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | INDUCTION: By heat shock, UVB, salt, wounding, ethylene and methyl jasmonate (PubMed:11162426, PubMed:12218065). Induced by infection with the fungal pathogen Golovinomyces cichoracearum (powdery mildew) and the bacterial pathogen Pseudomonas syringae pv tomato strain DC3000 (PubMed:22345509). {ECO:0000269|PubMed:11162426, ECO:0000269|PubMed:12218065, ECO:0000269|PubMed:22345509}. | |||||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_013636191.1 | 0.0 | PREDICTED: calmodulin-binding transcription activator 3 isoform X1 | ||||
| Swissprot | Q8GSA7 | 0.0 | CMTA3_ARATH; Calmodulin-binding transcription activator 3 | ||||
| TrEMBL | A0A3P6CF38 | 0.0 | A0A3P6CF38_BRAOL; Uncharacterized protein | ||||
| STRING | Bo4g149510.1 | 0.0 | (Brassica oleracea) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Malvids | OGEM8601 | 25 | 35 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT2G22300.2 | 0.0 | signal responsive 1 | ||||



