 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | GRMZM2G026643_P01 | ||||||||
| Common Name | Zm.224 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; Liliopsida; Petrosaviidae; commelinids; Poales; Poaceae; PACMAD clade; Panicoideae; Andropogonodae; Andropogoneae; Tripsacinae; Zea
|
||||||||
| Family | HD-ZIP | ||||||||
| Protein Properties | Length: 702aa MW: 75607 Da PI: 5.4629 | ||||||||
| Description | HD-ZIP family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | Homeobox | 70.3 | 2.4e-22 | 103 | 158 | 1 | 56 |
TT--SS--HHHHHHHHHHHHHSSS--HHHHHHHHHHCTS-HHHHHHHHHHHHHHHH CS
Homeobox 1 rrkRttftkeqleeLeelFeknrypsaeereeLAkklgLterqVkvWFqNrRakek 56
+++ +++t++q++eLe+lF+++++p++++r eL+k+lgL+ rqVk+WFqNrR+++k
GRMZM2G026643_P01 103 KKRYHRHTPQQIQELEALFKECPHPDEKQRGELSKRLGLDPRQVKFWFQNRRTQMK 158
678899***********************************************999 PP
| |||||||
| 2 | START | 186.8 | 1.1e-58 | 312 | 545 | 1 | 206 |
HHHHHHHHHHHHHHHHC-TT-EEEE......EXCCTTEEEEEEESSS.......SCEEEEEEEECCSCHHH.HHHHHHCCCGGCT-TT-S.. CS
START 1 elaeeaaqelvkkalaeepgWvkss......esengdevlqkfeeskv......dsgealrasgvvdmvla.llveellddkeqWdetla.. 77
ela +a++elvk+a+++ep+W s + +n +e+ +f++s + + +ea+r+sg+v+ + lve+l+d++ +W+ ++
GRMZM2G026643_P01 312 ELAISAMDELVKLAQVDEPLWLPSLsgspdkKLLNFEEYAHSFSPSVGavkpvgYVSEASRESGLVIIDNSlALVETLMDVR-RWSDMFScm 402
57889*****************999899988888999999999998779*****************9876658999999999.********* PP
..EEEEEEEECTT......EEEEEEEEXXTTXX-SSX.EEEEEEEEEEE.TTS-EEEEEEEEE..-TTS--....-TTSEE-EESSEEEEEE CS
START 78 ..kaetlevissg......galqlmvaelqalsplvp.RdfvfvRyirqlgagdwvivdvSvd..seqkppe...sssvvRaellpSgilie 155
ka++le ++sg gal lm+aelq+lsplvp R+++f+R+++ql +g w++vdvS+d ++++++ +++ +R+++lpSg++++
GRMZM2G026643_P01 403 iaKATVLEEVTSGiagsrnGALLLMKAELQVLSPLVPiREVTFLRFCKQLAEGAWAVVDVSIDglVRDHNSGtasNAGNIRCRRLPSGCVMQ 494
**************************************************************965566666677899*************** PP
EECTCEEEEEEEE-EE--SSXXHHHHHHHHHHHHHHHHHHHHHHTXXXXXX CS
START 156 pksnghskvtwvehvdlkgrlphwllrslvksglaegaktwvatlqrqcek 206
+++ng++kvtwve ++++++++h+l+r+l++sgla+ga++w+a lqrqce+
GRMZM2G026643_P01 495 DTPNGYCKVTWVEYTEYDEASVHQLYRPLIRSGLAFGARRWLAMLQRQCEC 545
*************************************************96 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| Gene3D | G3DSA:1.10.10.60 | 8.7E-24 | 88 | 160 | IPR009057 | Homeodomain-like |
| SuperFamily | SSF46689 | 8.77E-22 | 88 | 160 | IPR009057 | Homeodomain-like |
| PROSITE profile | PS50071 | 18.285 | 100 | 160 | IPR001356 | Homeobox domain |
| SMART | SM00389 | 2.8E-21 | 101 | 164 | IPR001356 | Homeobox domain |
| CDD | cd00086 | 3.38E-20 | 102 | 160 | No hit | No description |
| Pfam | PF00046 | 8.0E-20 | 103 | 158 | IPR001356 | Homeobox domain |
| PROSITE pattern | PS00027 | 0 | 135 | 158 | IPR017970 | Homeobox, conserved site |
| PROSITE profile | PS50848 | 47.244 | 303 | 548 | IPR002913 | START domain |
| SuperFamily | SSF55961 | 1.95E-31 | 305 | 545 | No hit | No description |
| CDD | cd08875 | 1.42E-115 | 307 | 544 | No hit | No description |
| SMART | SM00234 | 2.2E-45 | 312 | 545 | IPR002913 | START domain |
| Pfam | PF01852 | 2.5E-51 | 313 | 545 | IPR002913 | START domain |
| SuperFamily | SSF55961 | 5.86E-10 | 565 | 673 | No hit | No description |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0006355 | Biological Process | regulation of transcription, DNA-templated | ||||
| GO:0009827 | Biological Process | plant-type cell wall modification | ||||
| GO:0042335 | Biological Process | cuticle development | ||||
| GO:0043481 | Biological Process | anthocyanin accumulation in tissues in response to UV light | ||||
| GO:0048765 | Biological Process | root hair cell differentiation | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0008289 | Molecular Function | lipid binding | ||||
| GO:0043565 | Molecular Function | sequence-specific DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 702 aa Download sequence Send to blast |
MSFGSLFDGG SGGGGGGGGM QFPFSTGFSS SPALSLGLEN PGGGMVGRML PVGGAPAAGG 60 MARDAADAEN DSRSGSDHLD AMSAGAGAED EDDAEPGNPR KRKKRYHRHT PQQIQELEAL 120 FKECPHPDEK QRGELSKRLG LDPRQVKFWF QNRRTQMKTQ LERHENALLK QENDKLRAEN 180 MAIREAMRSP MCGSCGSPAM LGEVSLEEQH LCIENARLKD ELNRVYALAT KFLGKPMPVL 240 SGPMLQPNLS LPMPSSSLEL AVGGLRGLGS IPSLDEFAGG VSSPLGTVIT PARATGSAPP 300 PMVGVDRSML LELAISAMDE LVKLAQVDEP LWLPSLSGSP DKKLLNFEEY AHSFSPSVGA 360 VKPVGYVSEA SRESGLVIID NSLALVETLM DVRRWSDMFS CMIAKATVLE EVTSGIAGSR 420 NGALLLMKAE LQVLSPLVPI REVTFLRFCK QLAEGAWAVV DVSIDGLVRD HNSGTASNAG 480 NIRCRRLPSG CVMQDTPNGY CKVTWVEYTE YDEASVHQLY RPLIRSGLAF GARRWLAMLQ 540 RQCECLAILM SPDTVSANDS SVITQEGKRS MLKLARRMTE NFCAGVSASS AREWSKLDGA 600 AGSIGEDVRV MARKSVDEPG EPPGVVLSAA TSVWVPVAPE KLFNFLRDEQ LRAEWDILSN 660 GGPMQEMANI AKGQEHGNSV SLLRASHECQ PEQHADPPGD LH |
| Nucleic Localization Signal ? help Back to Top | |||
|---|---|---|---|
| No. | Start | End | Sequence |
| 1 | 98 | 104 | PRKRKKR |
| 2 | 99 | 103 | RKRKK |
| Expression -- UniGene ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| UniGene ID | E-value | Expressed in | ||||
| Zm.224 | 0.0 | ear| embryo| meristem| ovary| pericarp| shoot | ||||
| Expression -- Microarray ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | ID | |||||
| Expression Atlas | GRMZM2G026643 | |||||
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Probable transcription factor. {ECO:0000250}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
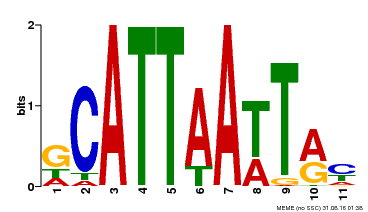
| Motif ID | Method | Source | Motif file |
| MP00421 | DAP | Transfer from AT4G00730 | Download |
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | GRMZM2G026643_P01 |
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | BT064453 | 0.0 | BT064453.1 Zea mays full-length cDNA clone ZM_BFc0170E20 mRNA, complete cds. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | NP_001105493.1 | 0.0 | outer cell layer 1 | ||||
| Swissprot | Q6EPF0 | 0.0 | ROC5_ORYSJ; Homeobox-leucine zipper protein ROC5 | ||||
| TrEMBL | C0P834 | 0.0 | C0P834_MAIZE; Homeobox-leucine zipper protein ANTHOCYANINLESS 2 | ||||
| STRING | GRMZM2G026643_P02 | 0.0 | (Zea mays) | ||||
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT4G00730.1 | 0.0 | HD-ZIP family protein | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | GRMZM2G026643_P01 |
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



