 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | FANhyb_rscf00000584.1.g00003.1 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; fabids; Rosales; Rosaceae; Rosoideae; Potentilleae; Fragariinae; Fragaria
|
||||||||
| Family | FAR1 | ||||||||
| Protein Properties | Length: 825aa MW: 95011.4 Da PI: 6.8148 | ||||||||
| Description | FAR1 family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | FAR1 | 88.5 | 9.6e-28 | 67 | 154 | 1 | 91 |
FAR1 1 kfYneYAkevGFsvrkskskkskrngeitkrtfvCskegkreeekkktekerrtraetrtgCkaklkvkkekdgkwevt 79
+fY+eYAk++GF++++++s++sk+++e+++++f Cs++g + e+++ ++r+ ++++t+Cka+++vk++ dgkw ++
FANhyb_rscf00000584.1.g00003.1 67 SFYQEYAKSMGFTTSIKNSRRSKKSKEFIDAKFACSRYGVTPESDSG---SSRRPTVKKTDCKASMHVKRRADGKWIIH 142
5****************************************999888...77888899********************* PP
FAR1 80 kleleHnHelap 91
++ +eHnHel p
FANhyb_rscf00000584.1.g00003.1 143 EFIKEHNHELLP 154
*********975 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| Pfam | PF03101 | 3.0E-25 | 67 | 154 | IPR004330 | FAR1 DNA binding domain |
| Pfam | PF10551 | 2.5E-24 | 274 | 367 | IPR018289 | MULE transposase domain |
| PROSITE profile | PS50966 | 10.225 | 554 | 590 | IPR007527 | Zinc finger, SWIM-type |
| Pfam | PF04434 | 5.9E-6 | 561 | 587 | IPR007527 | Zinc finger, SWIM-type |
| SMART | SM00575 | 1.3E-7 | 565 | 592 | IPR006564 | Zinc finger, PMZ-type |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0010018 | Biological Process | far-red light signaling pathway | ||||
| GO:0042753 | Biological Process | positive regulation of circadian rhythm | ||||
| GO:0045893 | Biological Process | positive regulation of transcription, DNA-templated | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| GO:0008270 | Molecular Function | zinc ion binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 825 aa Download sequence Send to blast |
MLVDGQDNMN DGRVIENMIV VPLVEEAQNS GGRVVSSPKR DVQLFEGDAD LEPKNGIEFD 60 SHEAAYSFYQ EYAKSMGFTT SIKNSRRSKK SKEFIDAKFA CSRYGVTPES DSGSSRRPTV 120 KKTDCKASMH VKRRADGKWI IHEFIKEHNH ELLPALAYHF RIHRNVKLAE KNNMDILQAV 180 SERTRKMYVE MSRQSGGYQN TGFLRADTKY QFDKCQDLGL DEGDAQVMLE YFKRIQKDNP 240 NFFYAIDLNE EQRVRNLFWV DAKSRSDYKS FNDVVSFDTS YIKINEKLPF APFVGVNHHY 300 QPMLLGCALI ADETKATFVW LLKTWVRAMG GLNPKVIISD QDNALKAAIE EVFQDARHCF 360 SLWNILEKIP EVLAPVVKRH ENFLPKFHKC IFKSWTDEQF DLKWFKMVTR FELQDDEWIR 420 SLYDDRKRWV PTYLGDTFLA GMCTTMRSES MNSFFDKYIH KKITLREFVK QYGTILQNRY 480 DEEAIADFDT WHKQPALKSP SPWEKQMSTI YTHAVFRKFQ VEVLGVVGCQ PKKEHEDGLT 540 TTFRVQDCEK EEYFMVTWTE SKSEVSCSCR LFEYKGFLCR HSLIVLQICG LSSIPPHYIL 600 KRWTKDAKSG QLILEGSELV QTRVQRYNDL CKRAIELSEE GSLSEETYNI ALRTLVEALK 660 NCVNVNNSNT TVVDFSSSVH GIREAEEENQ GSLAAKPSRK RNTNRKRKVQ AEQDVMIGEA 720 QDSLQQMDNL SSDGIALSGY YGAQHGLVQL NLMEPPHDNY YVNQESMQGL GQLNSIAPSD 780 DGFFGTQQIH GLGQLDFRPS SSFSYSLQDE PHLRSAELHG SSSRH |
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Transcription activator that recognizes and binds to the DNA consensus sequence 5'-CACGCGC-3'. Activates the expression of FHY1 and FHL involved in light responses. Positive regulator of chlorophyll biosynthesis via the activation of HEMB1 gene expression. {ECO:0000269|PubMed:11889039, ECO:0000269|PubMed:12753585, ECO:0000269|PubMed:18033885, ECO:0000269|PubMed:22634759}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00434 | DAP | Transfer from AT4G15090 | Download |
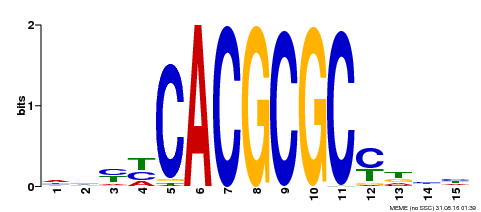
| |||
| Regulation -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | INDUCTION: Down-regulated after exposure to far-red light. Subject to a negative feedback regulation by PHYA signaling. {ECO:0000269|PubMed:18033885}. | |||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_004305896.1 | 0.0 | PREDICTED: protein FAR-RED IMPAIRED RESPONSE 1 | ||||
| Refseq | XP_011467678.1 | 0.0 | PREDICTED: protein FAR-RED IMPAIRED RESPONSE 1 | ||||
| Swissprot | Q9SWG3 | 0.0 | FAR1_ARATH; Protein FAR-RED IMPAIRED RESPONSE 1 | ||||
| TrEMBL | A0A2P6S0S3 | 0.0 | A0A2P6S0S3_ROSCH; Putative transcription factor FAR family | ||||
| STRING | XP_004305896.1 | 0.0 | (Fragaria vesca) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Fabids | OGEF40 | 30 | 530 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT4G15090.1 | 0.0 | FAR1 family protein | ||||



