 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | FANhyb_rscf00000230.1.g00005.1 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; fabids; Rosales; Rosaceae; Rosoideae; Potentilleae; Fragariinae; Fragaria
|
||||||||
| Family | MYB | ||||||||
| Protein Properties | Length: 479aa MW: 52359.6 Da PI: 5.5865 | ||||||||
| Description | MYB family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | Myb_DNA-binding | 38.4 | 2.8e-12 | 14 | 73 | 1 | 48 |
TSSS-HHHHHHHHHHHHHTTTT-HHHHHHHHT............TTS-HHHHHHHHHHHT CS
Myb_DNA-binding 1 rgrWTteEdellvdavkqlGggtWktIartmg............kgRtlkqcksrwqkyl 48
+grWT+eEd+ l+++++ G g+W++ ++ + R++k+c++rw +yl
FANhyb_rscf00000230.1.g00005.1 14 KGRWTAEEDQTLINYIRANGEGSWRSLPKNAAvqflqvqsidrgLLRCGKSCRLRWINYL 73
79*****************************99***********9*************97 PP
| |||||||
| 2 | Myb_DNA-binding | 51.5 | 2.3e-16 | 79 | 124 | 1 | 48 |
TSSS-HHHHHHHHHHHHHTTTT-HHHHHHHHTTTS-HHHHHHHHHHHT CS
Myb_DNA-binding 1 rgrWTteEdellvdavkqlGggtWktIartmgkgRtlkqcksrwqkyl 48
rg+ T++E+++++++++ lG++ W++Ia+ ++ gRt++++k++w+++l
FANhyb_rscf00000230.1.g00005.1 79 RGNITPQEEDIIIKLHASLGNR-WSLIASQLP-GRTDNEIKNYWNSHL 124
7999******************.*********.************986 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| Gene3D | G3DSA:1.10.10.60 | 4.8E-20 | 5 | 76 | IPR009057 | Homeodomain-like |
| PROSITE profile | PS51294 | 11.994 | 9 | 73 | IPR017930 | Myb domain |
| SuperFamily | SSF46689 | 1.56E-25 | 11 | 120 | IPR009057 | Homeodomain-like |
| SMART | SM00717 | 2.9E-9 | 13 | 75 | IPR001005 | SANT/Myb domain |
| Pfam | PF00249 | 3.0E-11 | 14 | 73 | IPR001005 | SANT/Myb domain |
| CDD | cd00167 | 8.90E-7 | 16 | 73 | No hit | No description |
| PROSITE profile | PS51294 | 25.801 | 74 | 128 | IPR017930 | Myb domain |
| Gene3D | G3DSA:1.10.10.60 | 2.3E-25 | 77 | 128 | IPR009057 | Homeodomain-like |
| SMART | SM00717 | 3.4E-16 | 78 | 126 | IPR001005 | SANT/Myb domain |
| Pfam | PF00249 | 8.6E-15 | 79 | 124 | IPR001005 | SANT/Myb domain |
| CDD | cd00167 | 7.71E-11 | 83 | 124 | No hit | No description |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0009651 | Biological Process | response to salt stress | ||||
| GO:0009723 | Biological Process | response to ethylene | ||||
| GO:0009733 | Biological Process | response to auxin | ||||
| GO:0009813 | Biological Process | flavonoid biosynthetic process | ||||
| GO:0045893 | Biological Process | positive regulation of transcription, DNA-templated | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 479 aa Download sequence Send to blast |
MGRAPCCEKV GLKKGRWTAE EDQTLINYIR ANGEGSWRSL PKNAAVQFLQ VQSIDRGLLR 60 CGKSCRLRWI NYLRADLKRG NITPQEEDII IKLHASLGNR WSLIASQLPG RTDNEIKNYW 120 NSHLSRKIDA FRRPINKIHM PSASAAAAAS SSSTTTTAAD EINMAMKLGP PSSKRRGRGG 180 RTSRWAMKKN KVTYASTAIN HHLDSCTHNH DDKSATMTSS SSPPPPSAAA AAVAAGTGQK 240 ETQALVGGGP SDGSAAGKLD DGGEVAFNDL LMDVANEIIS DPTGVLTLIG ERQSDDGDDD 300 TGVIRDDEDT GKLLCGPNKV MTNATAAASH EEEHHQLDQI SATNLSSSSN SNSKSSYGEA 360 EAAAFDRDDW YNSCSNYSPG SAAAATCKFD QQKQQRDGIN GTVNDEMRWD WESDIHRYDY 420 LWNVDQKDDE NMLSWLWEQQ AADDHHLDHS NKSRSEAVVV DDDCDPADKH NAILAWLLS |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 1a5j_A | 5e-23 | 12 | 129 | 5 | 109 | B-MYB |
| Search in ModeBase | ||||||
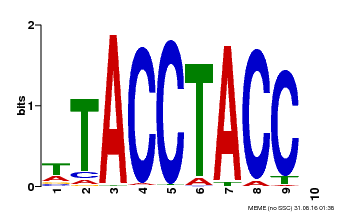
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00627 | PBM | Transfer from PK06182.1 | Download |
| |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_004301809.1 | 0.0 | PREDICTED: transcription factor MYB12-like | ||||
| TrEMBL | A0A2P6PIB4 | 0.0 | A0A2P6PIB4_ROSCH; Putative transcription factor MYB-HB-like family | ||||
| STRING | XP_004301809.1 | 0.0 | (Fragaria vesca) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Fabids | OGEF31 | 34 | 817 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT3G62610.1 | 3e-67 | myb domain protein 11 | ||||



