 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | FANhyb_rscf00000078.1.g00008.1 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; fabids; Rosales; Rosaceae; Rosoideae; Potentilleae; Fragariinae; Fragaria
|
||||||||
| Family | MYB_related | ||||||||
| Protein Properties | Length: 472aa MW: 52002 Da PI: 7.3803 | ||||||||
| Description | MYB_related family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | Myb_DNA-binding | 53.4 | 6.1e-17 | 79 | 123 | 1 | 47 |
TSSS-HHHHHHHHHHHHHTTTT-HHHHHHHHTTTS-HHHHHHHHHHH CS
Myb_DNA-binding 1 rgrWTteEdellvdavkqlGggtWktIartmgkgRtlkqcksrwqky 47
r +WT+eE++++++a k++G g W+ I +++g ++t+ q++s+ qk+
FANhyb_rscf00000078.1.g00008.1 79 REKWTEEEHQKFLEALKLYGRG-WRQIEEHVG-TKTAVQIRSHAQKF 123
789*******************.*********.************98 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| SuperFamily | SSF46689 | 1.05E-16 | 73 | 129 | IPR009057 | Homeodomain-like |
| PROSITE profile | PS51294 | 21.573 | 74 | 128 | IPR017930 | Myb domain |
| Gene3D | G3DSA:1.10.10.60 | 1.1E-9 | 76 | 126 | IPR009057 | Homeodomain-like |
| TIGRFAMs | TIGR01557 | 2.7E-16 | 77 | 126 | IPR006447 | Myb domain, plants |
| SMART | SM00717 | 2.8E-13 | 78 | 126 | IPR001005 | SANT/Myb domain |
| Pfam | PF00249 | 9.9E-15 | 79 | 122 | IPR001005 | SANT/Myb domain |
| CDD | cd00167 | 2.44E-11 | 81 | 124 | No hit | No description |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0007623 | Biological Process | circadian rhythm | ||||
| GO:0009651 | Biological Process | response to salt stress | ||||
| GO:0009723 | Biological Process | response to ethylene | ||||
| GO:0046686 | Biological Process | response to cadmium ion | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 472 aa Download sequence Send to blast |
MLVFGWLVIS YKFLQDYFAS ELNCDQSEAT GSNASLAVGN CSPKGDAQSK LVTELQDFSS 60 FGNDQALKVR KPYTITKQRE KWTEEEHQKF LEALKLYGRG WRQIEEHVGT KTAVQIRSHA 120 QKFFSKVAKE LPGTGEGSLK PIEIPPPRPK RKPMHPYPRK HNGSPERRSP SPNFSVAEKG 180 QESPTSVLSV PGSDIIGSPA LEQYNRSRPS TSCTTDMRSA NLSHVEKEND YATSSPSTEE 240 AKETMPHVDS TLDKPLSVVQ KDASASGCTS TTEGDAGRVG TSMSIKLFGR TVLVSDLHKQ 300 SSSGTKDCTS SIYMAKQENH ESESGKFGEP LQWDRSDTRL SLGGLDCNWN SLACGLTINR 360 VGQQDESPKS VPANSDAPPP LPWWVNLPFC QVVCVDSRVD ELSKQNERSC AGSNDGLVTS 420 IDSVCEEPRS VEPCNSRKGF VPYKRCLADT NRNSSVIVSE EKRDRQRARV CS |
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00146 | DAP | Transfer from AT1G18330 | Download |
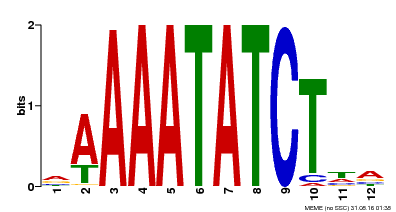
| |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_011464451.1 | 0.0 | PREDICTED: protein REVEILLE 7-like isoform X1 | ||||
| TrEMBL | A0A2P6P1K8 | 0.0 | A0A2P6P1K8_ROSCH; Putative transcription factor MYB-HB-like family | ||||
| STRING | XP_004299482.1 | 0.0 | (Fragaria vesca) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Fabids | OGEF7313 | 32 | 44 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT1G18330.1 | 3e-60 | MYB_related family protein | ||||



