 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | FANhyb_rscf00000065.1.g00001.1 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; fabids; Rosales; Rosaceae; Rosoideae; Potentilleae; Fragariinae; Fragaria
|
||||||||
| Family | MYB_related | ||||||||
| Protein Properties | Length: 355aa MW: 40107 Da PI: 9.8103 | ||||||||
| Description | MYB_related family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | Myb_DNA-binding | 44.3 | 4.1e-14 | 21 | 65 | 1 | 47 |
TSSS-HHHHHHHHHHHHHTTTT-HHHHHHHHTTTS-HHHHHHHHHHH CS
Myb_DNA-binding 1 rgrWTteEdellvdavkqlGggtWktIartmgkgRtlkqcksrwqky 47
r +WT+eE++++++a +++ + Wk+I +g +t q++s+ qky
FANhyb_rscf00000065.1.g00001.1 21 RESWTEEEHDKFLEALQLFDRD-WKKIEDFVG-SKTVIQIRSHAQKY 65
789*****************77.*********.*************9 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| SuperFamily | SSF46689 | 1.64E-17 | 15 | 71 | IPR009057 | Homeodomain-like |
| PROSITE profile | PS51294 | 16.291 | 16 | 70 | IPR017930 | Myb domain |
| TIGRFAMs | TIGR01557 | 1.2E-18 | 19 | 68 | IPR006447 | Myb domain, plants |
| Gene3D | G3DSA:1.10.10.60 | 3.6E-7 | 19 | 71 | IPR009057 | Homeodomain-like |
| SMART | SM00717 | 1.1E-11 | 20 | 68 | IPR001005 | SANT/Myb domain |
| Pfam | PF00249 | 9.2E-12 | 21 | 65 | IPR001005 | SANT/Myb domain |
| CDD | cd00167 | 1.09E-8 | 23 | 66 | No hit | No description |
| Pfam | PF14374 | 2.8E-12 | 231 | 278 | IPR025755 | 60S ribosomal protein L4, C-terminal domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0009651 | Biological Process | response to salt stress | ||||
| GO:0009723 | Biological Process | response to ethylene | ||||
| GO:0009733 | Biological Process | response to auxin | ||||
| GO:0009737 | Biological Process | response to abscisic acid | ||||
| GO:0009739 | Biological Process | response to gibberellin | ||||
| GO:0009751 | Biological Process | response to salicylic acid | ||||
| GO:0009753 | Biological Process | response to jasmonic acid | ||||
| GO:0032922 | Biological Process | circadian regulation of gene expression | ||||
| GO:0043966 | Biological Process | histone H3 acetylation | ||||
| GO:0046686 | Biological Process | response to cadmium ion | ||||
| GO:0048573 | Biological Process | photoperiodism, flowering | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 355 aa Download sequence Send to blast |
MSSSDGSGKK VRKPYTITKS RESWTEEEHD KFLEALQLFD RDWKKIEDFV GSKTVIQIRS 60 HAQKYFLKVQ KNGTTAHVPP PRPKRKASHP YPQKASKNEF IPFQASLAYP SSMNAIASGY 120 CPWDEASIFL NAGPDQIGVA RISKCDPSGI GGSSRTLPSS EITMQGSQAP VRHGLPDFAE 180 VYSFIGGVFD PDTQGHAQKL KEMDPINFET VLLVMRNLSI NLSSPDFEPI AVRPIKKDVK 240 RAPLKKNPLK NLNTMLRLNP YAKTAKRMAL LAEEHRLKSK KEKLEQKRKV TKNSEIMYVV 300 VLACVLVLEF EVYEAFEQEE ATKIKAAGKA WYQTMISDSD YTEFDVFQKW LGVSQ |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 4v3p_LD | 5e-23 | 232 | 355 | 274 | 372 | 60S ribosomal protein L4/L1 |
| 4v7e_CC | 7e-23 | 232 | 355 | 307 | 405 | 60S ribosomal protein L4 |
| Search in ModeBase | ||||||
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Transcriptional activator of evening element (EE)-containing clock-controlled genes. Forms a negative feedback loop with APRR5. Regulates the pattern of histone H3 acetylation of the TOC1 promoter. {ECO:0000269|PubMed:21205033, ECO:0000269|PubMed:21474993, ECO:0000269|PubMed:21483796, ECO:0000269|PubMed:23638299}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00338 | DAP | Transfer from AT3G09600 | Download |
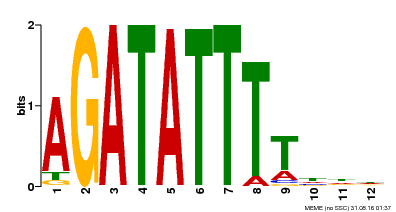
| |||
| Regulation -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | INDUCTION: Circadian-regulation. Peak of expression in the afternoon. Down-regulated by cold. {ECO:0000269|PubMed:21205033, ECO:0000269|PubMed:22902701, ECO:0000269|PubMed:23638299}. | |||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_004302837.1 | 1e-163 | PREDICTED: protein REVEILLE 8 | ||||
| Swissprot | Q8RWU3 | 1e-115 | RVE8_ARATH; Protein REVEILLE 8 | ||||
| TrEMBL | A0A2P6RAJ3 | 1e-150 | A0A2P6RAJ3_ROSCH; Putative transcription factor MYB-HB-like family | ||||
| STRING | XP_004302837.1 | 1e-162 | (Fragaria vesca) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Fabids | OGEF6735 | 34 | 49 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT3G09600.1 | 1e-117 | MYB_related family protein | ||||
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



