 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Eucgr.C04090.1.p | ||||||||
| Common Name | EUGRSUZ_C04090, LOC104438439 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; malvids; Myrtales; Myrtaceae; Myrtoideae; Eucalypteae; Eucalyptus
|
||||||||
| Family | FAR1 | ||||||||
| Protein Properties | Length: 825aa MW: 94820.8 Da PI: 7.1434 | ||||||||
| Description | FAR1 family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | FAR1 | 88.7 | 8e-28 | 61 | 148 | 1 | 91 |
FAR1 1 kfYneYAkevGFsvrkskskkskrngeitkrtfvCskegkreeekkktekerrtraetrtgCkaklkvkkekdgkwevtkleleHnHelap 91
+fY+eYAk++GF++++++s++sk+++e+++++f Cs++g + e+++ ++r+ ++++t+Cka+++vk+++dgkw +++++++HnHel p
Eucgr.C04090.1.p 61 SFYQEYAKSMGFTTSIKNSRRSKKSKEFIDAKFACSRYGVTPESESG---SSRRPSVKKTDCKASMHVKRRTDGKWIIHEFVKDHNHELLP 148
5***************************************9999877...77888899******************************975 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| Pfam | PF03101 | 2.2E-25 | 61 | 148 | IPR004330 | FAR1 DNA binding domain |
| Pfam | PF10551 | 1.9E-22 | 268 | 361 | IPR018289 | MULE transposase domain |
| PROSITE profile | PS50966 | 9.26 | 548 | 584 | IPR007527 | Zinc finger, SWIM-type |
| Pfam | PF04434 | 8.1E-5 | 554 | 581 | IPR007527 | Zinc finger, SWIM-type |
| SMART | SM00575 | 1.8E-6 | 559 | 586 | IPR006564 | Zinc finger, PMZ-type |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0010018 | Biological Process | far-red light signaling pathway | ||||
| GO:0042753 | Biological Process | positive regulation of circadian rhythm | ||||
| GO:0045893 | Biological Process | positive regulation of transcription, DNA-templated | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| GO:0008270 | Molecular Function | zinc ion binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 825 aa Download sequence Send to blast |
MDTRDGVQSG AGENMVSGTD EVQNRGSDFT PSETAVTGFE GDIDFEPRNG IEFDSHEAAY 60 SFYQEYAKSM GFTTSIKNSR RSKKSKEFID AKFACSRYGV TPESESGSSR RPSVKKTDCK 120 ASMHVKRRTD GKWIIHEFVK DHNHELLPAL AYHFRIHRNI KLAEKNNIDI LHAVSERTRK 180 MYVEMSRQSA GHPHACALHD KNDHQPDKGR SLTLDEGDGQ IILEYFKHIQ KENPSFFYAI 240 DLNEEQRLRN LLWVDAKSRN DYASFNDVVS LDTSYVKSND KFPFVPFVGV NHHSQTMLLG 300 CALIADKTRS TFVWLLKTWL RAMGGNSPSV ILSYQDKCLK EAIEEVLPQT RHCYLLWHIL 360 EKVHENLSHV INNHENFLSK FNKCIFKSWT DEQFDMRWWK MVSRFELQED EWVQMLYMER 420 KKWVPALLGE SFLAGMSTAQ RVDSMNSFFD RYIHKKITVK EFLRQYGVIL HNRCEEEAVA 480 DFDTWHKQPA LKSPSPWEKQ LSTVYTHAIF RKFQIEVLGV VACHPRKESE DGGTVIYRVE 540 DFEKKEYFMV TWNEVESEVS CFCHLFEYHG FLCRHAMVVL QISGFPSIPS RYIMKRWTKD 600 AKSRHSPDEG TKKMQTRVQR YNYLCRRAIE LSEEGSISEE AYAIAFRALV EALKNCVDVN 660 NSNNSMAECS TSICGLRETE SDNQGVTVNK TSKKRNTNKK RKVQEESDVL LAEAPDSLHE 720 MENLNTDGIT LNGYYGTQQN VQGLVQLNLM EPPHDAYYVS QPSIQGLGQL SSLAPSHDGF 780 FGTQPAMHVL GQVDFRTPST FNRSSPDQSH LRPTQVHGNS SRHP* |
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
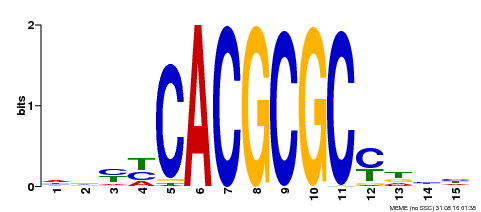
| UniProt | Transcription activator that recognizes and binds to the DNA consensus sequence 5'-CACGCGC-3'. Activates the expression of FHY1 and FHL involved in light responses. Positive regulator of chlorophyll biosynthesis via the activation of HEMB1 gene expression. {ECO:0000269|PubMed:11889039, ECO:0000269|PubMed:12753585, ECO:0000269|PubMed:18033885, ECO:0000269|PubMed:22634759}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00434 | DAP | Transfer from AT4G15090 | Download |
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | Eucgr.C04090.1.p |
| Regulation -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | INDUCTION: Down-regulated after exposure to far-red light. Subject to a negative feedback regulation by PHYA signaling. {ECO:0000269|PubMed:18033885}. | |||||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_010049888.1 | 0.0 | PREDICTED: protein FAR-RED IMPAIRED RESPONSE 1 isoform X1 | ||||
| Swissprot | Q9SWG3 | 0.0 | FAR1_ARATH; Protein FAR-RED IMPAIRED RESPONSE 1 | ||||
| TrEMBL | A0A059CW52 | 0.0 | A0A059CW52_EUCGR; Uncharacterized protein | ||||
| STRING | XP_010049888.1 | 0.0 | (Eucalyptus grandis) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Malvids | OGEM118 | 26 | 253 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT4G15090.1 | 0.0 | FAR1 family protein | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | Eucgr.C04090.1.p |
| Entrez Gene | 104438439 |



