 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Eucgr.A00553.1.p | ||||||||
| Common Name | EUGRSUZ_A00553 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; malvids; Myrtales; Myrtaceae; Myrtoideae; Eucalypteae; Eucalyptus
|
||||||||
| Family | C2H2 | ||||||||
| Protein Properties | Length: 454aa MW: 51348.3 Da PI: 8.5133 | ||||||||
| Description | C2H2 family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | zf-C2H2 | 11.1 | 0.0012 | 114 | 136 | 1 | 23 |
EEETTTTEEESSHHHHHHHHHHT CS
zf-C2H2 1 ykCpdCgksFsrksnLkrHirtH 23
y C++Cg + s ks+ ++Hi++H
Eucgr.A00553.1.p 114 YYCEYCGICRSKKSLITSHIQSH 136
89********************6 PP
| |||||||
| 2 | zf-C2H2 | 19 | 3.9e-06 | 160 | 181 | 2 | 23 |
EETTTTEEESSHHHHHHHHHHT CS
zf-C2H2 2 kCpdCgksFsrksnLkrHirtH 23
+C+ Cg +F+ + +L++H++ H
Eucgr.A00553.1.p 160 TCEECGATFKKPAYLRQHMQGH 181
6******************988 PP
| |||||||
| 3 | zf-C2H2 | 15.1 | 6.7e-05 | 187 | 211 | 1 | 23 |
EEET..TTTEEESSHHHHHHHHHHT CS
zf-C2H2 1 ykCp..dCgksFsrksnLkrHirtH 23
+kC+ dC+ s++rk++L rH H
Eucgr.A00553.1.p 187 FKCSieDCPASYRRKDHLARHFLQH 211
89*******************7766 PP
| |||||||
| 4 | zf-C2H2 | 20 | 1.9e-06 | 216 | 241 | 1 | 23 |
EEET..TTTEEESSHHHHHHHHHH.T CS
zf-C2H2 1 ykCp..dCgksFsrksnLkrHirt.H 23
++Cp C+ksF + n+krH++ H
Eucgr.A00553.1.p 216 FSCPvdGCNKSFAYQGNMKRHLNEiH 241
89********************9988 PP
| |||||||
| 5 | zf-C2H2 | 20.2 | 1.6e-06 | 258 | 282 | 1 | 23 |
EEET..TTTEEESSHHHHHHHHHHT CS
zf-C2H2 1 ykCp..dCgksFsrksnLkrHirtH 23
y+Cp Cgk+F+ s L++H +H
Eucgr.A00553.1.p 258 YVCPevGCGKVFKFASKLRKHEDSH 282
9********************9888 PP
| |||||||
| 6 | zf-C2H2 | 16.7 | 2.1e-05 | 348 | 373 | 1 | 23 |
EEET..TTTEEESSHHHHHHHHHH.T CS
zf-C2H2 1 ykCp..dCgksFsrksnLkrHirt.H 23
++C C +Fs+ksnL++H + H
Eucgr.A00553.1.p 348 FRCDveGCLSTFSTKSNLRQHVKAvH 373
899999***************99877 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| SMART | SM00355 | 18 | 114 | 136 | IPR015880 | Zinc finger, C2H2-like |
| PROSITE profile | PS50157 | 8.663 | 114 | 142 | IPR007087 | Zinc finger, C2H2 |
| PROSITE profile | PS50157 | 13.006 | 159 | 186 | IPR007087 | Zinc finger, C2H2 |
| SMART | SM00355 | 0.012 | 159 | 181 | IPR015880 | Zinc finger, C2H2-like |
| SuperFamily | SSF57667 | 2.7E-10 | 160 | 211 | No hit | No description |
| Gene3D | G3DSA:3.30.160.60 | 2.9E-8 | 160 | 185 | IPR013087 | Zinc finger C2H2-type/integrase DNA-binding domain |
| PROSITE pattern | PS00028 | 0 | 161 | 181 | IPR007087 | Zinc finger, C2H2 |
| Gene3D | G3DSA:3.30.160.60 | 3.6E-9 | 186 | 213 | IPR013087 | Zinc finger C2H2-type/integrase DNA-binding domain |
| PROSITE profile | PS50157 | 9.411 | 187 | 214 | IPR007087 | Zinc finger, C2H2 |
| SMART | SM00355 | 0.0069 | 187 | 211 | IPR015880 | Zinc finger, C2H2-like |
| PROSITE pattern | PS00028 | 0 | 189 | 211 | IPR007087 | Zinc finger, C2H2 |
| SuperFamily | SSF57667 | 1.35E-8 | 197 | 253 | No hit | No description |
| Gene3D | G3DSA:3.30.160.60 | 9.4E-8 | 214 | 242 | IPR013087 | Zinc finger C2H2-type/integrase DNA-binding domain |
| SMART | SM00355 | 1.6E-4 | 216 | 241 | IPR015880 | Zinc finger, C2H2-like |
| PROSITE profile | PS50157 | 13.069 | 216 | 246 | IPR007087 | Zinc finger, C2H2 |
| PROSITE pattern | PS00028 | 0 | 218 | 241 | IPR007087 | Zinc finger, C2H2 |
| SuperFamily | SSF57667 | 1.29E-5 | 256 | 310 | No hit | No description |
| Gene3D | G3DSA:3.30.160.60 | 6.1E-7 | 256 | 282 | IPR013087 | Zinc finger C2H2-type/integrase DNA-binding domain |
| PROSITE profile | PS50157 | 11.032 | 258 | 287 | IPR007087 | Zinc finger, C2H2 |
| SMART | SM00355 | 0.0026 | 258 | 282 | IPR015880 | Zinc finger, C2H2-like |
| PROSITE pattern | PS00028 | 0 | 260 | 282 | IPR007087 | Zinc finger, C2H2 |
| PROSITE profile | PS50157 | 8.995 | 290 | 313 | IPR007087 | Zinc finger, C2H2 |
| SMART | SM00355 | 1.5 | 290 | 313 | IPR015880 | Zinc finger, C2H2-like |
| PROSITE pattern | PS00028 | 0 | 292 | 313 | IPR007087 | Zinc finger, C2H2 |
| SMART | SM00355 | 29 | 316 | 337 | IPR015880 | Zinc finger, C2H2-like |
| SuperFamily | SSF57667 | 3.0E-9 | 329 | 390 | No hit | No description |
| Gene3D | G3DSA:3.30.160.60 | 2.8E-7 | 329 | 370 | IPR013087 | Zinc finger C2H2-type/integrase DNA-binding domain |
| PROSITE profile | PS50157 | 12.009 | 348 | 378 | IPR007087 | Zinc finger, C2H2 |
| SMART | SM00355 | 8.2E-4 | 348 | 373 | IPR015880 | Zinc finger, C2H2-like |
| PROSITE pattern | PS00028 | 0 | 350 | 373 | IPR007087 | Zinc finger, C2H2 |
| SMART | SM00355 | 27 | 379 | 405 | IPR015880 | Zinc finger, C2H2-like |
| PROSITE pattern | PS00028 | 0 | 381 | 405 | IPR007087 | Zinc finger, C2H2 |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0005730 | Cellular Component | nucleolus | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| GO:0008097 | Molecular Function | 5S rRNA binding | ||||
| GO:0046872 | Molecular Function | metal ion binding | ||||
| GO:0080084 | Molecular Function | 5S rDNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 454 aa Download sequence Send to blast |
MIRFVLDGSR FYPSYMLTLR LGLDVGNDSE VKPLPVRGSN LVPVGVYWAG LNLGPSRSRA 60 QNDTDPYDES SVISSNCLGF EVCSATSARA GARAMGGEES CEKPGPVFRD IRRYYCEYCG 120 ICRSKKSLIT SHIQSHHQDE VEAAMAQGVE GKEGAKANNT CEECGATFKK PAYLRQHMQG 180 HSLERPFKCS IEDCPASYRR KDHLARHFLQ HKGKIFSCPV DGCNKSFAYQ GNMKRHLNEI 240 HNDEEPSTSG ECKSLKQYVC PEVGCGKVFK FASKLRKHED SHVKLETVEA FCCECMKYFS 300 NVDYLKEHMR SAHQLVNCDT CGTKQLRKNI QRHLRTHEKK NSADSEAFRC DVEGCLSTFS 360 TKSNLRQHVK AVHLEVRPFV CGHKDCGKKF AYKHVRDHHE KTGCHVYTPG DFEELDEQFR 420 SRPRGGRKRK CPTIESLLRK RVTLPNELAF LSD* |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 2i13_A | 3e-18 | 161 | 402 | 19 | 190 | Aart |
| 2i13_B | 3e-18 | 161 | 402 | 19 | 190 | Aart |
| Search in ModeBase | ||||||
| Nucleic Localization Signal ? help Back to Top | |||
|---|---|---|---|
| No. | Start | End | Sequence |
| 1 | 421 | 429 | RPRGGRKRK |
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Essential protein (PubMed:22353599). Isoform 1 is a transcription activator the binds both 5S rDNA and 5S rRNA and stimulates the transcription of 5S rRNA gene (PubMed:12711688, PubMed:22353599). Isoform 1 regulates 5S rRNA levels during development (PubMed:22353599). {ECO:0000269|PubMed:12711688, ECO:0000269|PubMed:22353599}. | |||||
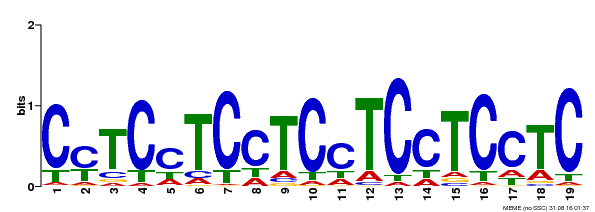
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00229 | DAP | Transfer from AT1G72050 | Download |
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | Eucgr.A00553.1.p |
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_010034379.1 | 0.0 | PREDICTED: transcription factor IIIA | ||||
| Swissprot | Q84MZ4 | 1e-131 | TF3A_ARATH; Transcription factor IIIA | ||||
| TrEMBL | A0A059DCN0 | 0.0 | A0A059DCN0_EUCGR; Uncharacterized protein | ||||
| STRING | XP_010034379.1 | 0.0 | (Eucalyptus grandis) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Malvids | OGEM4714 | 28 | 52 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT1G72050.2 | 1e-133 | transcription factor IIIA | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | Eucgr.A00553.1.p |



