 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | EcC035387.20 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; malvids; Myrtales; Myrtaceae; Myrtoideae; Eucalypteae; Eucalyptus
|
||||||||
| Family | MYB_related | ||||||||
| Protein Properties | Length: 483aa MW: 53141.6 Da PI: 6.5172 | ||||||||
| Description | MYB_related family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | Myb_DNA-binding | 53.9 | 4e-17 | 59 | 103 | 1 | 47 |
TSSS-HHHHHHHHHHHHHTTTT-HHHHHHHHTTTS-HHHHHHHHHHH CS
Myb_DNA-binding 1 rgrWTteEdellvdavkqlGggtWktIartmgkgRtlkqcksrwqky 47
r +WT eE+e++++a k++G g W+ I +++g ++t+ q++s+ qk+
EcC035387.20 59 REKWTDEEHERFLEALKLYGRG-WRQIEEHVG-TKTAVQIRSHAQKF 103
789*******************.*********.************98 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| SuperFamily | SSF46689 | 9.72E-17 | 53 | 107 | IPR009057 | Homeodomain-like |
| PROSITE profile | PS51294 | 21.548 | 54 | 108 | IPR017930 | Myb domain |
| TIGRFAMs | TIGR01557 | 1.9E-16 | 57 | 106 | IPR006447 | Myb domain, plants |
| SMART | SM00717 | 1.1E-12 | 58 | 106 | IPR001005 | SANT/Myb domain |
| Pfam | PF00249 | 2.8E-14 | 59 | 102 | IPR001005 | SANT/Myb domain |
| Gene3D | G3DSA:1.10.10.60 | 5.9E-9 | 59 | 99 | IPR009057 | Homeodomain-like |
| CDD | cd00167 | 1.19E-10 | 61 | 104 | No hit | No description |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0007623 | Biological Process | circadian rhythm | ||||
| GO:0009651 | Biological Process | response to salt stress | ||||
| GO:0009723 | Biological Process | response to ethylene | ||||
| GO:0046686 | Biological Process | response to cadmium ion | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 483 aa Download sequence Send to blast |
MAGEAQNEVA VSSALFTVPN RPSKVGSQLE AVDNLKELQV LENDQTPKVR KPYTISKQRE 60 KWTDEEHERF LEALKLYGRG WRQIEEHVGT KTAVQIRSHA QKFFSKVARG VSGSSEGVIK 120 PIEIPPPRPK RKPMHPYPRK SVDSKEVKLS YQQERSPSPI SSVADENTGS PTSVLSAHGS 180 DMLGSASLHQ QNRCSSPTSC TTDVPSIGLA VIEKQPEIFK EEDKGSLSST QMPSGLTLGN 240 LPSMNCDLSS KNTSSPEEIG GTEAPSIKLF GRTVLVTEIQ KPDHQEAGIK KSPAQKTGED 300 DSVTEDEGLG KPLSSVEFDM QLSLGLACGN NGFLHSGAHV ATAMKLQKED VESSRFTSDC 360 FLPWWPLYQG PPFIYVTPSD STLIQGPLDS HEDRKVEVEE IIKETSSETN TVAKKTEGYG 420 DNNVENNADG VFPKCQKHLE RRTHPGDCKK GFVPYKRCLA ERDVKSPVIV SEERQAQRTR 480 VCS |
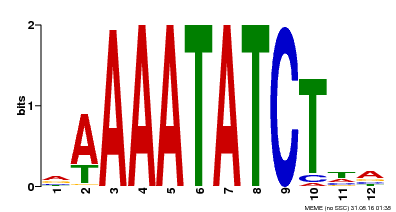
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00146 | DAP | Transfer from AT1G18330 | Download |
| |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_010037006.1 | 0.0 | PREDICTED: protein REVEILLE 7 | ||||
| Refseq | XP_010037007.1 | 0.0 | PREDICTED: protein REVEILLE 7 | ||||
| TrEMBL | A0A059A496 | 0.0 | A0A059A496_EUCGR; Uncharacterized protein | ||||
| STRING | XP_010037005.1 | 0.0 | (Eucalyptus grandis) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Malvids | OGEM9174 | 27 | 38 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT1G18330.1 | 7e-49 | MYB_related family protein | ||||



