 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | EMT08925 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; Liliopsida; Petrosaviidae; commelinids; Poales; Poaceae; BOP clade; Pooideae; Triticodae; Triticeae; Triticinae; Aegilops
|
||||||||
| Family | B3 | ||||||||
| Protein Properties | Length: 907aa MW: 104194 Da PI: 6.9927 | ||||||||
| Description | B3 family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | B3 | 51.9 | 1.4e-16 | 35 | 123 | 1 | 98 |
EEEE-..-HHHHTT-EE--HHH.HTT---..--SEEEEEETTS-EEEEEE..EEETTEEEE-TTHHHHHHHHT--TT-EEEEEE-SSSEE..EEEEE- CS
B3 1 ffkvltpsdvlksgrlvlpkkfaeehggkkeesktltledesgrsWevkliyrkksgryvltkGWkeFvkangLkegDfvvFkldgrsefelvvkvfr 98
ffk + ++++g +++p kf+++ gg s ++le+++g++++v + k + vl++GW++F++ +L+egD + Fk++g+s f +v +++
EMT08925 35 FFKLM--LGNFRNG-VTIPEKFVRSVGGM--ISELVKLETPDGNTYNVHI--AKELNNLVLRSGWSKFASVYELQEGDLLRFKYNGDSHF--KVEIYD 123
56666..6677776.***********776..6678***************..**********************************9999..888776 PP
| |||||||
| 2 | B3 | 43.9 | 4.1e-14 | 264 | 345 | 17 | 99 |
E--HHH.HTT---..--SEEEEEE.TTS-EEEEEE..EEETTEEEE-TTHHHHHHHHT--TT-EEEEEE-SS.SEE..EEEEE-S CS
B3 17 vlpkkfaeehggkkeesktltled.esgrsWevkliyrkksgryvltkGWkeFvkangLkegDfvvFkldgr.sefelvvkvfrk 99
+ k+ a +h ++ +tl + +++++W ++ + +++y l++GW +Fv++n+L+ gD++vF++ +r ++f++ v+++++
EMT08925 264 LICKDDAIKHLPR--KDEFITLCHaSHSKTWGAHYN-INADDTYHLSAGWLRFVDDNQLQKGDTCVFEVLKRqRSFTMAVHLLKA 345
5667777777322..3334555554899*******3.44445699***********************88777*******99876 PP
| |||||||
| 3 | B3 | 33.7 | 6.5e-11 | 436 | 500 | 35 | 99 |
EEEEEE.TTS-EEEEEE..EEETTEEEE-TTHHHHHHHHT--TT-EEEEEE-SS.SEE..EEEEE-S CS
B3 35 tltled.esgrsWevkliyrkksgryvltkGWkeFvkangLkegDfvvFkldgr.sefelvvkvfrk 99
+l ++ W+ ++ r ++r +l kGW++Fv +n+L+ D+++F+++++ ++ +++v+++rk
EMT08925 436 IFRLHRpGESAPWKAEF--RSFGSRRWLVKGWEQFVIDNELELDDVCLFERIENkKKLRMMVHIIRK 500
45555523447899999..5555555699***********************999**********98 PP
| |||||||
| 4 | B3 | 61.1 | 1.9e-19 | 557 | 648 | 1 | 99 |
EEEE-..-HHHHTT-EE--HHH.HTT---..--SEEEEEETTS-EEEEEE..EEETTEEEE-TTHHHHHHHHT--TT-EEEEEE-SSSEE..EEEEE-S CS
B3 1 ffkvltpsdvlksgrlvlpkkfaeehggkkeesktltledesgrsWevkliyrkksgryvltkGWkeFvkangLkegDfvvFkldgrsefelvvkvfrk 99
ffk + + +++r+++p kfa+++ ++++s+ le +sg++W v + +k + ++++GW +Fvka++L+e+D +vF+++g+s+f v +f++
EMT08925 557 FFKLM-T--QDSQQRFRIPDKFASSFIRQTQSSQGFDLEAPSGETWHVGV--TKVANDLFFSSGWGDFVKAHELQENDLLVFTFSGNSSF--EVLIFDA 648
56655.3..3455679**********777788899***************..**********************************9999..8888876 PP
| |||||||
| 5 | B3 | 52.2 | 1.1e-16 | 803 | 877 | 11 | 87 |
HHTT-EE--HHH.HTT.---..--SEEEEEE.TTS-EEEEEE..EEETTEEEE-TTHHHHHHHHT--TT-EEEEEE-SS CS
B3 11 lksgrlvlpkkfaeeh.ggkkeesktltled.esgrsWevkliyrkksgryvltkGWkeFvkangLkegDfvvFkldgr 87
+k++ lv+p +f++eh +s +++l + ++W v++ y+++s+r + ++ W +Fv++n+L+egD++vF+l++
EMT08925 803 HKNNFLVIPSEFVDEHlH-M--RSHEVVLLRpNREERWHVRY-YQGSSSRGFRGQPWAKFVRENELREGDVCVFELIKG 877
567889**********52.3..3445555552778*******.999999989999********************9965 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| SuperFamily | SSF101936 | 1.59E-19 | 28 | 127 | IPR015300 | DNA-binding pseudobarrel domain |
| Gene3D | G3DSA:2.40.330.10 | 3.4E-20 | 29 | 127 | IPR015300 | DNA-binding pseudobarrel domain |
| PROSITE profile | PS50863 | 12.478 | 32 | 125 | IPR003340 | B3 DNA binding domain |
| CDD | cd10017 | 7.32E-17 | 33 | 123 | No hit | No description |
| SMART | SM01019 | 7.6E-14 | 35 | 125 | IPR003340 | B3 DNA binding domain |
| Pfam | PF02362 | 2.0E-14 | 35 | 123 | IPR003340 | B3 DNA binding domain |
| Gene3D | G3DSA:2.40.330.10 | 3.4E-15 | 235 | 346 | IPR015300 | DNA-binding pseudobarrel domain |
| SuperFamily | SSF101936 | 3.53E-15 | 236 | 345 | IPR015300 | DNA-binding pseudobarrel domain |
| CDD | cd10017 | 1.03E-9 | 240 | 344 | No hit | No description |
| SMART | SM01019 | 0.0029 | 242 | 346 | IPR003340 | B3 DNA binding domain |
| Pfam | PF02362 | 2.9E-11 | 264 | 345 | IPR003340 | B3 DNA binding domain |
| PROSITE profile | PS50863 | 11.702 | 281 | 346 | IPR003340 | B3 DNA binding domain |
| Gene3D | G3DSA:2.40.330.10 | 1.7E-15 | 395 | 500 | IPR015300 | DNA-binding pseudobarrel domain |
| SuperFamily | SSF101936 | 4.32E-19 | 396 | 500 | IPR015300 | DNA-binding pseudobarrel domain |
| CDD | cd10017 | 1.96E-12 | 401 | 499 | No hit | No description |
| SMART | SM01019 | 6.4E-11 | 403 | 501 | IPR003340 | B3 DNA binding domain |
| PROSITE profile | PS50863 | 11.152 | 404 | 501 | IPR003340 | B3 DNA binding domain |
| Pfam | PF02362 | 4.4E-9 | 417 | 500 | IPR003340 | B3 DNA binding domain |
| SuperFamily | SSF101936 | 9.02E-22 | 550 | 651 | IPR015300 | DNA-binding pseudobarrel domain |
| Gene3D | G3DSA:2.40.330.10 | 7.3E-22 | 554 | 651 | IPR015300 | DNA-binding pseudobarrel domain |
| PROSITE profile | PS50863 | 14.663 | 554 | 649 | IPR003340 | B3 DNA binding domain |
| CDD | cd10017 | 3.02E-20 | 557 | 647 | No hit | No description |
| SMART | SM01019 | 2.7E-16 | 557 | 649 | IPR003340 | B3 DNA binding domain |
| Pfam | PF02362 | 1.7E-16 | 557 | 648 | IPR003340 | B3 DNA binding domain |
| Gene3D | G3DSA:2.40.330.10 | 3.2E-19 | 784 | 878 | IPR015300 | DNA-binding pseudobarrel domain |
| SuperFamily | SSF101936 | 1.16E-18 | 785 | 877 | IPR015300 | DNA-binding pseudobarrel domain |
| CDD | cd10017 | 2.56E-15 | 790 | 877 | No hit | No description |
| SMART | SM01019 | 6.1E-13 | 792 | 899 | IPR003340 | B3 DNA binding domain |
| PROSITE profile | PS50863 | 13.239 | 793 | 897 | IPR003340 | B3 DNA binding domain |
| Pfam | PF02362 | 2.5E-14 | 794 | 877 | IPR003340 | B3 DNA binding domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0006355 | Biological Process | regulation of transcription, DNA-templated | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0005773 | Cellular Component | vacuole | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 907 aa Download sequence Send to blast |
MPVQSDQRMD KSCTRCKDYV AHQYWDHMDN RKKRFFKLML GNFRNGVTIP EKFVRSVGGM 60 ISELVKLETP DGNTYNVHIA KELNNLVLRS GWSKFASVYE LQEGDLLRFK YNGDSHFKVE 120 IYDPSACEKE IYCVVMNHNP GLQKRSVPHD NQMLSPEGER LAKRQNGCCS DGCKTSEMNP 180 AGSPHKPTKK EVPSSEDARS SGERKTFTRS HYFLATGCNL TDAHKVEVDK IEQNIRPEIP 240 LYVKTMSSAS LVDGFLIICN RSYLICKDDA IKHLPRKDEF ITLCHASHSK TWGAHYNINA 300 DDTYHLSAGW LRFVDDNQLQ KGDTCVFEVL KRQRSFTMAV HLLKASHHQP PGFPTSSNSL 360 RPEVKSFRPE DKVRLSRFTT LEGLLKTKVY EKVEAIKPEI PVFVSIMMKT NVSSRTPSLA 420 FSLDYAKYFL PGENQIFRLH RPGESAPWKA EFRSFGSRRW LVKGWEQFVI DNELELDDVC 480 LFERIENKKK LRMMVHIIRK EDIKILQIFV GGHILIKCTN SGNPTKYEMM GEKGCESCKK 540 WQEHYYWEHM DVSKTKFFKL MTQDSQQRFR IPDKFASSFI RQTQSSQGFD LEAPSGETWH 600 VGVTKVANDL FFSSGWGDFV KAHELQENDL LVFTFSGNSS FEVLIFDATG CEKLSSLFAG 660 AGMHKHFDGM VGQQVEQYSP SDDSDEDDDN DDDDDDDDDD TSVPSQLIES PHKVSTLRKF 720 SGKTKPSKEL RESPNSSSSC DVKHEETEEE ESDDDTYADF DYCYSRAAKQ LPDDEKSEII 780 GLASIQPGNP AFMTVLLRAH LQHKNNFLVI PSEFVDEHLH MRSHEVVLLR PNREERWHVR 840 YYQGSSSRGF RGQPWAKFVR ENELREGDVC VFELIKGARN EKKLARTTMA IHVARRRKSN 900 GRFALVD |
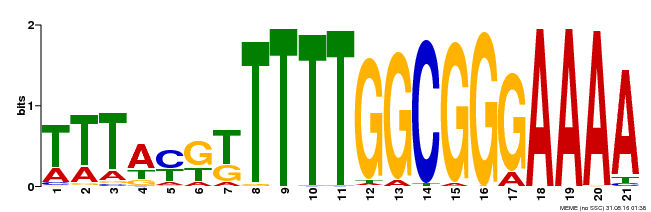
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00468 | DAP | Transfer from AT4G33280 | Download |
| |||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | AK373848 | 0.0 | AK373848.1 Hordeum vulgare subsp. vulgare mRNA for predicted protein, partial cds, clone: NIASHv3045G04. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_024318846.1 | 0.0 | LOW QUALITY PROTEIN: B3 domain-containing protein LOC_Os12g40080 | ||||
| TrEMBL | M8B5P7 | 0.0 | M8B5P7_AEGTA; B3 domain-containing protein | ||||
| STRING | EMT08925 | 0.0 | (Aegilops tauschii) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Monocots | OGMP562 | 34 | 179 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT4G33280.1 | 7e-31 | B3 family protein | ||||



