 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Do014240.1 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; Liliopsida; Petrosaviidae; commelinids; Poales; Poaceae; PACMAD clade; Panicoideae; Panicodae; Paniceae; Dichantheliinae; Dichanthelium
|
||||||||
| Family | C2H2 | ||||||||
| Protein Properties | Length: 596aa MW: 67969.3 Da PI: 7.5565 | ||||||||
| Description | C2H2 family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | zf-C2H2 | 13.7 | 0.00018 | 347 | 368 | 2 | 23 |
EETTTTEEESSHHHHHHHHHHT CS
zf-C2H2 2 kCpdCgksFsrksnLkrHirtH 23
+C Cg +F + +Lk+H+++H
Do014240.1 347 RCLECGACFQKPAHLKQHMQSH 368
7*******************99 PP
| |||||||
| 2 | zf-C2H2 | 24 | 1e-07 | 374 | 398 | 1 | 23 |
EEET..TTTEEESSHHHHHHHHHHT CS
zf-C2H2 1 ykCp..dCgksFsrksnLkrHirtH 23
++Cp dC+ s++rk++L+rH+ +H
Do014240.1 374 FVCPleDCPFSYKRKDHLNRHMLKH 398
89*********************99 PP
| |||||||
| 3 | zf-C2H2 | 11.9 | 0.00071 | 405 | 428 | 3 | 23 |
ET..TTTEEESSHHHHHHHHHH.T CS
zf-C2H2 3 Cp..dCgksFsrksnLkrHirt.H 23
C+ C+++Fs k n++rH + H
Do014240.1 405 CTvdGCDRRFSIKANMQRHVKEfH 428
88889***************9988 PP
| |||||||
| 4 | zf-C2H2 | 15.8 | 3.9e-05 | 440 | 464 | 1 | 23 |
EEET..TTTEEESSHHHHHHHHHHT CS
zf-C2H2 1 ykCp..dCgksFsrksnLkrHirtH 23
+ C+ C+k F+ s Lk+H +H
Do014240.1 440 FICKeeGCNKAFKYLSKLKKHEESH 464
789999***************9988 PP
| |||||||
| 5 | zf-C2H2 | 17.6 | 1.1e-05 | 531 | 555 | 2 | 23 |
EET..TTTEEESSHHHHHHHHHH.T CS
zf-C2H2 2 kCp..dCgksFsrksnLkrHirt.H 23
kC+ C +sFs+ksnL++H++ H
Do014240.1 531 KCTfeGCARSFSNKSNLTTHMKAcH 555
789999****************955 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| PROSITE profile | PS51257 | 5 | 1 | 20 | No hit | No description |
| PROSITE profile | PS51375 | 8.769 | 44 | 74 | IPR002885 | Pentatricopeptide repeat |
| TIGRFAMs | TIGR00756 | 4.1E-4 | 49 | 72 | IPR002885 | Pentatricopeptide repeat |
| Pfam | PF01535 | 4.0E-5 | 49 | 72 | IPR002885 | Pentatricopeptide repeat |
| Gene3D | G3DSA:1.25.40.10 | 3.5E-4 | 50 | 105 | IPR011990 | Tetratricopeptide-like helical domain |
| Pfam | PF13041 | 1.2E-10 | 74 | 122 | IPR002885 | Pentatricopeptide repeat |
| PROSITE profile | PS51375 | 12.014 | 75 | 109 | IPR002885 | Pentatricopeptide repeat |
| TIGRFAMs | TIGR00756 | 7.2E-7 | 77 | 111 | IPR002885 | Pentatricopeptide repeat |
| PROSITE profile | PS51375 | 8.429 | 110 | 140 | IPR002885 | Pentatricopeptide repeat |
| PROSITE profile | PS51375 | 6.807 | 146 | 176 | IPR002885 | Pentatricopeptide repeat |
| Pfam | PF01535 | 0.011 | 149 | 173 | IPR002885 | Pentatricopeptide repeat |
| Gene3D | G3DSA:1.25.40.10 | 3.5E-4 | 209 | 241 | IPR011990 | Tetratricopeptide-like helical domain |
| PROSITE profile | PS51375 | 7.618 | 212 | 246 | IPR002885 | Pentatricopeptide repeat |
| SMART | SM00355 | 22 | 287 | 309 | IPR015880 | Zinc finger, C2H2-like |
| SMART | SM00355 | 0.094 | 346 | 368 | IPR015880 | Zinc finger, C2H2-like |
| PROSITE profile | PS50157 | 12.03 | 346 | 373 | IPR007087 | Zinc finger, C2H2 |
| Gene3D | G3DSA:3.30.160.60 | 9.5E-4 | 347 | 364 | IPR013087 | Zinc finger C2H2-type/integrase DNA-binding domain |
| PROSITE pattern | PS00028 | 0 | 348 | 368 | IPR007087 | Zinc finger, C2H2 |
| SuperFamily | SSF57667 | 9.24E-11 | 358 | 400 | No hit | No description |
| Gene3D | G3DSA:3.30.160.60 | 7.3E-12 | 365 | 400 | IPR013087 | Zinc finger C2H2-type/integrase DNA-binding domain |
| Pfam | PF00096 | 1.5E-5 | 374 | 398 | IPR007087 | Zinc finger, C2H2 |
| SMART | SM00355 | 1.6E-4 | 374 | 398 | IPR015880 | Zinc finger, C2H2-like |
| PROSITE profile | PS50157 | 9.141 | 374 | 403 | IPR007087 | Zinc finger, C2H2 |
| PROSITE pattern | PS00028 | 0 | 376 | 398 | IPR007087 | Zinc finger, C2H2 |
| SuperFamily | SSF57667 | 1.88E-7 | 384 | 433 | No hit | No description |
| Gene3D | G3DSA:3.30.160.60 | 2.1E-8 | 401 | 430 | IPR013087 | Zinc finger C2H2-type/integrase DNA-binding domain |
| SMART | SM00355 | 0.027 | 403 | 428 | IPR015880 | Zinc finger, C2H2-like |
| PROSITE profile | PS50157 | 10.512 | 403 | 433 | IPR007087 | Zinc finger, C2H2 |
| PROSITE pattern | PS00028 | 0 | 405 | 428 | IPR007087 | Zinc finger, C2H2 |
| Gene3D | G3DSA:3.30.160.60 | 5.3E-7 | 436 | 464 | IPR013087 | Zinc finger C2H2-type/integrase DNA-binding domain |
| PROSITE profile | PS50157 | 10.201 | 440 | 464 | IPR007087 | Zinc finger, C2H2 |
| SMART | SM00355 | 0.0087 | 440 | 464 | IPR015880 | Zinc finger, C2H2-like |
| PROSITE pattern | PS00028 | 0 | 442 | 464 | IPR007087 | Zinc finger, C2H2 |
| SMART | SM00355 | 8.2 | 472 | 497 | IPR015880 | Zinc finger, C2H2-like |
| PROSITE pattern | PS00028 | 0 | 474 | 497 | IPR007087 | Zinc finger, C2H2 |
| SMART | SM00355 | 18 | 500 | 521 | IPR015880 | Zinc finger, C2H2-like |
| SuperFamily | SSF57667 | 7.33E-9 | 515 | 572 | No hit | No description |
| SMART | SM00355 | 0.0066 | 530 | 555 | IPR015880 | Zinc finger, C2H2-like |
| PROSITE profile | PS50157 | 12.383 | 530 | 560 | IPR007087 | Zinc finger, C2H2 |
| Gene3D | G3DSA:3.30.160.60 | 3.7E-7 | 531 | 556 | IPR013087 | Zinc finger C2H2-type/integrase DNA-binding domain |
| PROSITE pattern | PS00028 | 0 | 532 | 555 | IPR007087 | Zinc finger, C2H2 |
| Gene3D | G3DSA:3.30.160.60 | 7.2E-5 | 559 | 582 | IPR013087 | Zinc finger C2H2-type/integrase DNA-binding domain |
| SMART | SM00355 | 5.8 | 561 | 587 | IPR015880 | Zinc finger, C2H2-like |
| PROSITE profile | PS50157 | 8.787 | 561 | 587 | IPR007087 | Zinc finger, C2H2 |
| PROSITE pattern | PS00028 | 0 | 563 | 587 | IPR007087 | Zinc finger, C2H2 |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0005730 | Cellular Component | nucleolus | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| GO:0005515 | Molecular Function | protein binding | ||||
| GO:0008097 | Molecular Function | 5S rRNA binding | ||||
| GO:0046872 | Molecular Function | metal ion binding | ||||
| GO:0080084 | Molecular Function | 5S rDNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 596 aa Download sequence Send to blast |
MVMEGHRPTH VTIVGVLSAC ADIGALDLGR VIHGYGSKCN ASTNIIVTNA LIDMYAKSGR 60 IEMAFSLFEE VQLKDAFTWT TMISSFTVQG DGRKALELFW DMLRSGVVPN SVTFISVLSA 120 CSHAGLIEEG TELFGTMREI YNIDPQLEHY GCMIDLLGRG GLLEEAEALI ADMNVEPDIV 180 MWRSILSACL VHRNDRLAEI AGKEIIKREP GDDGVYVILW NMYASSNKWR EAREMRQQML 240 TRKIFKKPGC SWIEVDGVVH EFLMCSGDEI DGDASVGQKS FKDIRRYKCE FCTVVRSKKC 300 LIQAHMVAHH KCVLFKVFPK IYLQDELSKS EIYNSDGEKI VCEEENRCLE CGACFQKPAH 360 LKQHMQSHSN ERLFVCPLED CPFSYKRKDH LNRHMLKHQG KLFGCTVDGC DRRFSIKANM 420 QRHVKEFHED ENVTKGNQQF ICKEEGCNKA FKYLSKLKKH EESHVKLNYM EVVCCEPGCM 480 KMFTNVECLR AHNQSCHQHV QCEICGEKHL KKNIKRHLQA HDEFPSGERM KCTFEGCARS 540 FSNKSNLTTH MKACHDQLKP FTCRVAGCGK AFTYKHVRDN HENSSAHVYI EVSPTT |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 5iww_D | 2e-33 | 1 | 244 | 12 | 335 | PLS9-PPR |
| Search in ModeBase | ||||||
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Essential protein (PubMed:22353599). Isoform 1 is a transcription activator the binds both 5S rDNA and 5S rRNA and stimulates the transcription of 5S rRNA gene (PubMed:12711688, PubMed:22353599). Isoform 1 regulates 5S rRNA levels during development (PubMed:22353599). {ECO:0000269|PubMed:12711688, ECO:0000269|PubMed:22353599}. | |||||
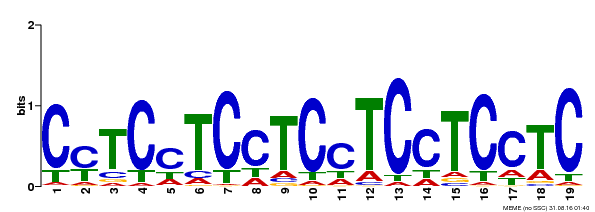
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00229 | DAP | Transfer from AT1G72050 | Download |
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | Do014240.1 |
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | - | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_025795605.1 | 0.0 | transcription factor IIIA-like isoform X1 | ||||
| Swissprot | Q84MZ4 | 1e-98 | TF3A_ARATH; Transcription factor IIIA | ||||
| TrEMBL | K3Z0X6 | 0.0 | K3Z0X6_SETIT; Uncharacterized protein | ||||
| STRING | Si020193m | 0.0 | (Setaria italica) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Monocots | OGMP2382 | 38 | 82 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT1G72050.2 | 1e-101 | transcription factor IIIA | ||||



