 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Do010218.1 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; Liliopsida; Petrosaviidae; commelinids; Poales; Poaceae; PACMAD clade; Panicoideae; Panicodae; Paniceae; Dichantheliinae; Dichanthelium
|
||||||||
| Family | G2-like | ||||||||
| Protein Properties | Length: 482aa MW: 51226.1 Da PI: 5.5742 | ||||||||
| Description | G2-like family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | G2-like | 100.4 | 1.2e-31 | 268 | 323 | 1 | 56 |
G2-like 1 kprlrWtpeLHerFveaveqLGGsekAtPktilelmkvkgLtlehvkSHLQkYRla 56
k+r++W+peLH+rFv+a++ LGG+++AtPk+i+elmkv+gLt+++vkSHLQkYRl+
Do010218.1 268 KARRCWSPELHRRFVNALQILGGAQVATPKQIRELMKVDGLTNDEVKSHLQKYRLH 323
68****************************************************97 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| SuperFamily | SSF46689 | 2.12E-17 | 265 | 326 | IPR009057 | Homeodomain-like |
| PROSITE profile | PS51294 | 13.69 | 265 | 325 | IPR017930 | Myb domain |
| Gene3D | G3DSA:1.10.10.60 | 3.4E-28 | 266 | 326 | IPR009057 | Homeodomain-like |
| TIGRFAMs | TIGR01557 | 2.8E-26 | 268 | 323 | IPR006447 | Myb domain, plants |
| Pfam | PF00249 | 1.0E-6 | 270 | 321 | IPR001005 | SANT/Myb domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0009266 | Biological Process | response to temperature stimulus | ||||
| GO:0009740 | Biological Process | gibberellic acid mediated signaling pathway | ||||
| GO:0010452 | Biological Process | histone H3-K36 methylation | ||||
| GO:0048579 | Biological Process | negative regulation of long-day photoperiodism, flowering | ||||
| GO:1903507 | Biological Process | negative regulation of nucleic acid-templated transcription | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0043565 | Molecular Function | sequence-specific DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 482 aa Download sequence Send to blast |
MESSPSDLTL DYKPNGNGNG GGAYAVIPKQ QEPLVDGHHL TTEQTTQKLR EFLARLEEER 60 LKIDAFKREL PLCMHLLNHA MEAYRQQLEA YQMGSLQGAP ARPLVLEEFI PLKNMNLGID 120 TAADKMGNPP SEKASWMESA QLWNGPAATA AVDTVAKGPQ TPKESSEHPL PTDTLGALDA 180 TSAGQRNGGA FLPFAKDKAA AEAAALPELA LAPAEKDAAE ADRKPPYLDA AGTNGGLGAR 240 RDVQNGVKPA SNAPDGQAPP PQPQTHRKAR RCWSPELHRR FVNALQILGG AQVATPKQIR 300 ELMKVDGLTN DEVKSHLQKY RLHTRRPMPP ASTAATAAPQ LVVLGGIWVP PEYATQAAGQ 360 AIYGAHPATQ PHYTAVAAAQ EYYPPHAAVH HLQHHPAAMV HRAAAAPPPP QAAYKVAMAG 420 SPPESSEGRG SAGGGSGGGR ERSESIEEED EGEEREEDDD DDEMAAAKGD GEESAGAGAI 480 KY |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 6j4k_A | 3e-14 | 268 | 321 | 2 | 55 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j4k_B | 3e-14 | 268 | 321 | 2 | 55 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j4r_A | 3e-14 | 268 | 321 | 1 | 54 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j4r_B | 3e-14 | 268 | 321 | 1 | 54 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j4r_C | 3e-14 | 268 | 321 | 1 | 54 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j4r_D | 3e-14 | 268 | 321 | 1 | 54 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j5b_A | 3e-14 | 268 | 321 | 2 | 55 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j5b_C | 3e-14 | 268 | 321 | 2 | 55 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j5b_D | 3e-14 | 268 | 321 | 2 | 55 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j5b_F | 3e-14 | 268 | 321 | 2 | 55 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j5b_H | 3e-14 | 268 | 321 | 2 | 55 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j5b_J | 3e-14 | 268 | 321 | 2 | 55 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| Search in ModeBase | ||||||
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Probable transcription factor that may play a role in regulatory networks controlling development and metabolism. {ECO:0000250|UniProtKB:Q6Z869}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00261 | DAP | Transfer from AT2G03500 | Download |
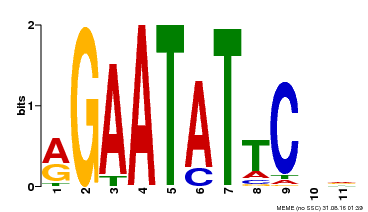
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | Do010218.1 |
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | BT086572 | 1e-175 | BT086572.2 Zea mays full-length cDNA clone ZM_BFc0184N23 mRNA, complete cds. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_025819732.1 | 0.0 | transcription factor NIGTH1 | ||||
| Swissprot | Q5VRW2 | 1e-142 | NOH1_ORYSJ; Transcription factor NIGTH1 | ||||
| TrEMBL | A0A1E5VUK6 | 0.0 | A0A1E5VUK6_9POAL; Myb family transcription factor EFM | ||||
| STRING | Si001230m | 0.0 | (Setaria italica) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Monocots | OGMP9773 | 33 | 38 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT1G68670.1 | 4e-47 | G2-like family protein | ||||
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



