 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Do006873.1 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; Liliopsida; Petrosaviidae; commelinids; Poales; Poaceae; PACMAD clade; Panicoideae; Panicodae; Paniceae; Dichantheliinae; Dichanthelium
|
||||||||
| Family | HD-ZIP | ||||||||
| Protein Properties | Length: 822aa MW: 89246 Da PI: 6.0135 | ||||||||
| Description | HD-ZIP family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | Homeobox | 58 | 1.7e-18 | 5 | 63 | 3 | 57 |
--SS--HHHHHHHHHHHHHSSS--HHHHHHHHHHC....TS-HHHHHHHHHHHHHHHHC CS
Homeobox 3 kRttftkeqleeLeelFeknrypsaeereeLAkkl....gLterqVkvWFqNrRakekk 57
k ++t+eq+e+Le+l++++++ps +r++L + + +++ +q+kvWFqNrR +ek+
Do006873.1 5 KYVRYTPEQVEALERLYYECPKPSSLRRQQLVRDCpvlaNVDPKQIKVWFQNRRCREKQ 63
6789*****************************************************97 PP
| |||||||
| 2 | START | 89.6 | 6.7e-29 | 153 | 279 | 2 | 126 |
HHHHHHHHHHHHHHHC-TT-EEEEEXCCTTEEEEEEESSS.SCEEEEEEEECCSCHHHHHHHHHCCCGGCT-TT-SEEEEEEEECTT..EEEEEEEEXX CS
START 2 laeeaaqelvkkalaeepgWvkssesengdevlqkfeeskvdsgealrasgvvdmvlallveellddkeqWdetlakaetlevissg..galqlmvael 98
+aee+++e+++ka+ ++ Wv+++ +++g++++ +++ s+++ g a+ra+g+v m++a v+e+l+d++ W ++++++e+++v+ g g+++l +++l
Do006873.1 153 IAEETLTEFLSKATGTAVEWVQMPGMKPGPDSIGIIAISHGCAGVAARACGLVGMEPA-KVAEILKDRPLWLRDCRSMEVVNVLPAGnnGTIELLYMQL 250
789*******************************************************.8888888888******************9*********** PP
TTXX-SSX.EEEEEEEEEEE.TTS-EEEE CS
START 99 qalsplvp.RdfvfvRyirqlgagdwviv 126
+a+ +l+p Rdf+ +Ry+ l +g++v+
Do006873.1 251 YAPMTLAPaRDFWLLRYTSILDDGSLVVS 279
********************999999986 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| PROSITE profile | PS50071 | 15.402 | 1 | 64 | IPR001356 | Homeobox domain |
| SMART | SM00389 | 1.2E-14 | 2 | 68 | IPR001356 | Homeobox domain |
| CDD | cd00086 | 1.92E-15 | 5 | 65 | No hit | No description |
| Pfam | PF00046 | 3.9E-16 | 5 | 63 | IPR001356 | Homeobox domain |
| SuperFamily | SSF46689 | 2.95E-16 | 6 | 66 | IPR009057 | Homeodomain-like |
| Gene3D | G3DSA:1.10.10.60 | 1.2E-17 | 7 | 63 | IPR009057 | Homeodomain-like |
| CDD | cd14686 | 3.99E-6 | 57 | 96 | No hit | No description |
| PROSITE profile | PS50848 | 13.087 | 143 | 278 | IPR002913 | START domain |
| SuperFamily | SSF55961 | 7.28E-21 | 144 | 284 | No hit | No description |
| Gene3D | G3DSA:3.30.530.20 | 1.8E-10 | 150 | 280 | IPR023393 | START-like domain |
| SMART | SM00234 | 7.1E-8 | 152 | 315 | IPR002913 | START domain |
| Pfam | PF01852 | 4.6E-26 | 153 | 279 | IPR002913 | START domain |
| Pfam | PF08670 | 3.2E-42 | 647 | 818 | IPR013978 | MEKHLA |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0009965 | Biological Process | leaf morphogenesis | ||||
| GO:0010014 | Biological Process | meristem initiation | ||||
| GO:0010075 | Biological Process | regulation of meristem growth | ||||
| GO:0010087 | Biological Process | phloem or xylem histogenesis | ||||
| GO:0048263 | Biological Process | determination of dorsal identity | ||||
| GO:0080060 | Biological Process | integument development | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| GO:0008289 | Molecular Function | lipid binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 822 aa Download sequence Send to blast |
MDASKYVRYT PEQVEALERL YYECPKPSSL RRQQLVRDCP VLANVDPKQI KVWFQNRRCR 60 EKQRKESSRL QALNRKLTAM NKLLMEENDR LQKQVSQLVY ENGYYRQQTQ SAGLATTDTS 120 CESVVTSGQQ NVVGAGQPQA QPRDASPAGL MSIAEETLTE FLSKATGTAV EWVQMPGMKP 180 GPDSIGIIAI SHGCAGVAAR ACGLVGMEPA KVAEILKDRP LWLRDCRSME VVNVLPAGNN 240 GTIELLYMQL YAPMTLAPAR DFWLLRYTSI LDDGSLVVSF YAGNIYSIIV RPLYESSAMV 300 AQKMSMAALR YLRQVAHEDT HSVITGWGRQ PTALRALSQK LTRGFNEALN GLADDGWSVI 360 ESDGVDDVCV SVNSSPSKVI NCNAAFNNGL PIVSSSVLCA KASMLLQDVS PPALLRFMRE 420 QRSQWADSNL DALFASAMKP NFCNLPMSRL GGFSGQVILP LAHTFDPEEF LEVIKLGNAS 480 TYQDALMHRD LFLLQMYNGV DENTVGTCSE LLFAPIDASF SDDSPLLPSG FRIIPIDSPL 540 DTSSPNCTLD LASTLEVGTP RSRIPGGGGS GKAACAGSKA VMTIAFQFAF ESHLQDSVAT 600 MARQYMRSII ASVQRIALAL SSSRLVPQVG GVSHAPAAAV ASPEAATLSR WICQSYRFHF 660 GAELIKSADA SECETGLKAL WHHASAILCC SLKVCSENCF GVARSGLLSS GLTQDYTSSM 720 QAMPVFTFAN QSGLDMLETT LVALQDITLE KVLGDXGRKN LCAELPGVVE QGFACIPGGL 780 CVSGLGRPVS YEKALAWKVL DDDSAAHCIC FMFVNWSFVS SM |
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Probable transcription factor. {ECO:0000250}. | |||||
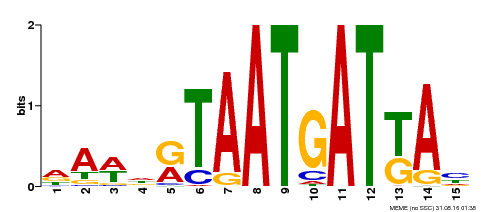
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00200 | DAP | Transfer from AT1G52150 | Download |
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | Do006873.1 |
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_025816436.1 | 0.0 | homeobox-leucine zipper protein HOX29-like | ||||
| Swissprot | A2WLR5 | 0.0 | HOX29_ORYSI; Homeobox-leucine zipper protein HOX29 | ||||
| TrEMBL | A0A1E5V3I0 | 0.0 | A0A1E5V3I0_9POAL; Homeobox-leucine zipper protein HOX29 | ||||
| STRING | Pavir.Eb00113.1.p | 0.0 | (Panicum virgatum) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Monocots | OGMP374 | 38 | 197 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT1G52150.1 | 0.0 | HD-ZIP family protein | ||||



