 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | DCAR_030989 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; asterids; campanulids; Apiales; Apiineae; Apiaceae; Apioideae; Scandiceae; Daucinae; Daucus
|
||||||||
| Family | C2H2 | ||||||||
| Protein Properties | Length: 432aa MW: 49620.3 Da PI: 8.2626 | ||||||||
| Description | C2H2 family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | zf-C2H2 | 11.7 | 0.00077 | 20 | 42 | 1 | 23 |
EEETTTTEEESSHHHHHHHHHHT CS
zf-C2H2 1 ykCpdCgksFsrksnLkrHirtH 23
y C++Cg + s k +L +Hi +H
DCAR_030989 20 YYCEYCGICRSKKTLLSSHILSH 42
89*******************96 PP
| |||||||
| 2 | zf-C2H2 | 18.8 | 4.4e-06 | 65 | 86 | 2 | 23 |
EETTTTEEESSHHHHHHHHHHT CS
zf-C2H2 2 kCpdCgksFsrksnLkrHirtH 23
+C+ Cg sF + +Lk+H+++H
DCAR_030989 65 TCEECGLSFQKPAHLKQHMQSH 86
6*******************99 PP
| |||||||
| 3 | zf-C2H2 | 19 | 3.9e-06 | 92 | 116 | 1 | 23 |
EEET..TTTEEESSHHHHHHHHHHT CS
zf-C2H2 1 ykCp..dCgksFsrksnLkrHirtH 23
+ Cp dC+ s++rk++L+rH+ H
DCAR_030989 92 FMCPvdDCDSSYRRKDHLNRHLLQH 116
79*******************9887 PP
| |||||||
| 4 | zf-C2H2 | 11.1 | 0.0013 | 121 | 146 | 1 | 23 |
EEET..TTTEEESSHHHHHHHHHH.T CS
zf-C2H2 1 ykCp..dCgksFsrksnLkrHirt.H 23
++C+ C+ +Fs +sn+krH + H
DCAR_030989 121 FECSsmGCSHRFSIQSNMKRHVKEfH 146
89999*****************9988 PP
| |||||||
| 5 | zf-C2H2 | 19.9 | 2e-06 | 218 | 242 | 1 | 23 |
EEET..TTTEEESSHHHHHHHHHHT CS
zf-C2H2 1 ykCp..dCgksFsrksnLkrHirtH 23
y+Cp Cgk+F+ s L++H +H
DCAR_030989 218 YTCPepGCGKVFKYASRLRKHEDSH 242
99*******************9888 PP
| |||||||
| 6 | zf-C2H2 | 18.5 | 5.5e-06 | 309 | 333 | 2 | 23 |
EET..TTTEEESSHHHHHHHHHH.T CS
zf-C2H2 2 kCp..dCgksFsrksnLkrHirt.H 23
kC+ dC +Fs+ksnL++H++ H
DCAR_030989 309 KCSfeDCLHTFSNKSNLRQHMKAvH 333
79999****************9888 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| SMART | SM00355 | 16 | 20 | 42 | IPR015880 | Zinc finger, C2H2-like |
| SuperFamily | SSF57667 | 7.32E-11 | 63 | 116 | No hit | No description |
| SMART | SM00355 | 0.0071 | 64 | 86 | IPR015880 | Zinc finger, C2H2-like |
| PROSITE profile | PS50157 | 12.986 | 64 | 91 | IPR007087 | Zinc finger, C2H2 |
| Gene3D | G3DSA:3.30.160.60 | 5.6E-8 | 65 | 92 | IPR013087 | Zinc finger C2H2-type/integrase DNA-binding domain |
| PROSITE pattern | PS00028 | 0 | 66 | 86 | IPR007087 | Zinc finger, C2H2 |
| SMART | SM00355 | 0.0021 | 92 | 116 | IPR015880 | Zinc finger, C2H2-like |
| PROSITE profile | PS50157 | 9.702 | 92 | 121 | IPR007087 | Zinc finger, C2H2 |
| Gene3D | G3DSA:3.30.160.60 | 7.9E-10 | 93 | 118 | IPR013087 | Zinc finger C2H2-type/integrase DNA-binding domain |
| PROSITE pattern | PS00028 | 0 | 94 | 116 | IPR007087 | Zinc finger, C2H2 |
| SuperFamily | SSF57667 | 7.42E-6 | 119 | 147 | No hit | No description |
| Gene3D | G3DSA:3.30.160.60 | 1.1E-6 | 119 | 148 | IPR013087 | Zinc finger C2H2-type/integrase DNA-binding domain |
| SMART | SM00355 | 0.88 | 121 | 146 | IPR015880 | Zinc finger, C2H2-like |
| PROSITE profile | PS50157 | 11.406 | 121 | 151 | IPR007087 | Zinc finger, C2H2 |
| PROSITE pattern | PS00028 | 0 | 123 | 146 | IPR007087 | Zinc finger, C2H2 |
| SMART | SM00355 | 0.022 | 179 | 204 | IPR015880 | Zinc finger, C2H2-like |
| PROSITE profile | PS50157 | 11.24 | 179 | 209 | IPR007087 | Zinc finger, C2H2 |
| Gene3D | G3DSA:3.30.160.60 | 1.3E-7 | 179 | 205 | IPR013087 | Zinc finger C2H2-type/integrase DNA-binding domain |
| PROSITE pattern | PS00028 | 0 | 181 | 204 | IPR007087 | Zinc finger, C2H2 |
| SuperFamily | SSF57667 | 5.41E-5 | 216 | 244 | No hit | No description |
| Gene3D | G3DSA:3.30.160.60 | 1.9E-7 | 216 | 242 | IPR013087 | Zinc finger C2H2-type/integrase DNA-binding domain |
| SMART | SM00355 | 2.2E-4 | 218 | 242 | IPR015880 | Zinc finger, C2H2-like |
| PROSITE profile | PS50157 | 11.468 | 218 | 247 | IPR007087 | Zinc finger, C2H2 |
| PROSITE pattern | PS00028 | 0 | 220 | 242 | IPR007087 | Zinc finger, C2H2 |
| SMART | SM00355 | 4.1 | 250 | 275 | IPR015880 | Zinc finger, C2H2-like |
| PROSITE pattern | PS00028 | 0 | 252 | 275 | IPR007087 | Zinc finger, C2H2 |
| SMART | SM00355 | 16 | 278 | 299 | IPR015880 | Zinc finger, C2H2-like |
| Gene3D | G3DSA:3.30.160.60 | 1.2E-5 | 289 | 330 | IPR013087 | Zinc finger C2H2-type/integrase DNA-binding domain |
| SuperFamily | SSF57667 | 1.15E-7 | 289 | 346 | No hit | No description |
| SMART | SM00355 | 8.0E-4 | 308 | 333 | IPR015880 | Zinc finger, C2H2-like |
| PROSITE profile | PS50157 | 11.655 | 308 | 338 | IPR007087 | Zinc finger, C2H2 |
| PROSITE pattern | PS00028 | 0 | 310 | 333 | IPR007087 | Zinc finger, C2H2 |
| SMART | SM00355 | 11 | 339 | 365 | IPR015880 | Zinc finger, C2H2-like |
| PROSITE pattern | PS00028 | 0 | 341 | 365 | IPR007087 | Zinc finger, C2H2 |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0005730 | Cellular Component | nucleolus | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| GO:0008097 | Molecular Function | 5S rRNA binding | ||||
| GO:0046872 | Molecular Function | metal ion binding | ||||
| GO:0080084 | Molecular Function | 5S rDNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 432 aa Download sequence Send to blast |
MEEGGTDITK GPVFKDIRRY YCEYCGICRS KKTLLSSHIL SHHQDEVNKR KADDNAEKEE 60 LKSNTCEECG LSFQKPAHLK QHMQSHLLER PFMCPVDDCD SSYRRKDHLN RHLLQHQGKL 120 FECSSMGCSH RFSIQSNMKR HVKEFHNDSS PIEVDALCSS SYRRKDLLNR HLQDQGELFE 180 CSVTGCCQKF SSQSNMKKHV KVFHDDSSPI EVDALKEYTC PEPGCGKVFK YASRLRKHED 240 SHVKLESTEA FCSDPGCMKY FSNEQCLKAH IQSCHSKVTC EVCGSKHLKK NLKRHLLTHK 300 TKPPSDRIKC SFEDCLHTFS NKSNLRQHMK AVHFEQKAFL CSIPGCGMSF TFKHVRDNHE 360 KSGCHLYVQG DFLETDEQFR SRPRGGRKRV CPSVETFTRK RVVPPSVGSD SLPDQGPNYM 420 SWLLSSENEE I* |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 1tf6_A | 5e-17 | 68 | 202 | 19 | 158 | PROTEIN (TRANSCRIPTION FACTOR IIIA) |
| 1tf6_D | 5e-17 | 68 | 202 | 19 | 158 | PROTEIN (TRANSCRIPTION FACTOR IIIA) |
| Search in ModeBase | ||||||
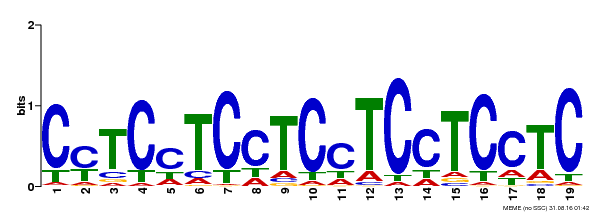
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00229 | DAP | Transfer from AT1G72050 | Download |
| |||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_017223703.1 | 0.0 | PREDICTED: transcription factor IIIA-like | ||||
| TrEMBL | A0A175YJJ2 | 0.0 | A0A175YJJ2_DAUCS; Uncharacterized protein | ||||
| STRING | VIT_08s0040g00240.t01 | 1e-153 | (Vitis vinifera) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Asterids | OGEA3644 | 24 | 47 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT1G72050.2 | 1e-132 | transcription factor IIIA | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | DCAR_030989 |



