 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | DCAR_028858 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; asterids; campanulids; Apiales; Apiineae; Apiaceae; Apioideae; Scandiceae; Daucinae; Daucus
|
||||||||
| Family | Trihelix | ||||||||
| Protein Properties | Length: 282aa MW: 33292.6 Da PI: 8.4573 | ||||||||
| Description | Trihelix family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | trihelix | 80.1 | 3.1e-25 | 32 | 114 | 1 | 86 |
trihelix 1 rWtkqevlaLiearremeerlrrgklkkplWeevskkmrergferspkqCkekwenlnkrykkikegekkrtsessstcpyfdqle 86
+W+ qe+++ + +r e++ ++ ++k++k lWe ++ km+e+g++rs++qCk+kw+nl +ryk ++++e ++ ++++py+++l+
DCAR_028858 32 QWSMQETRDFLMIRAELDPTFMETKRNKLLWELIATKMKEKGYNRSAEQCKCKWKNLVTRYKGCETIEPEG---MKQQFPYYNELQ 114
7********************************************************************97...45579*****98 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| SMART | SM00717 | 0.007 | 29 | 91 | IPR001005 | SANT/Myb domain |
| CDD | cd12203 | 3.90E-25 | 31 | 96 | No hit | No description |
| Pfam | PF13837 | 7.6E-21 | 32 | 116 | No hit | No description |
| PROSITE profile | PS50090 | 7.422 | 33 | 89 | IPR017877 | Myb-like domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0005730 | Cellular Component | nucleolus | ||||
| GO:0016592 | Cellular Component | mediator complex | ||||
| GO:0042802 | Molecular Function | identical protein binding | ||||
| GO:0043565 | Molecular Function | sequence-specific DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 282 aa Download sequence Send to blast |
MDAGHHHHLL HHQLQQQQVS VNIDTGGDRF PQWSMQETRD FLMIRAELDP TFMETKRNKL 60 LWELIATKMK EKGYNRSAEQ CKCKWKNLVT RYKGCETIEP EGMKQQFPYY NELQAIFAAR 120 MQRMLWNEAE GSGAGGSKKK ATQLSSDDED ENEESDGEQR GTTSGKKKRK VKQNPGSSNA 180 IGGNVNANVG GLKEMLEDFM RQQGEMEMRW LKTYEAREEE RRMREMEWRQ RMEELEKERI 240 MMERRWRERE EQRSAREEAR AEKRDALINA LLIKLRREDK S* |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 2ebi_A | 6e-13 | 29 | 112 | 3 | 86 | DNA binding protein GT-1 |
| Search in ModeBase | ||||||
| Nucleic Localization Signal ? help Back to Top | |||
|---|---|---|---|
| No. | Start | End | Sequence |
| 1 | 165 | 170 | KKKRKV |
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Probable transcription factor that binds specifically to the core DNA sequence 5'-GTTAC-3'. {ECO:0000269|PubMed:15044016}. | |||||
| UniProt | Probable transcription factor that may play a role in the induction of CAM4 in response to pathogen and salt. {ECO:0000269|PubMed:15310827}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00481 | DAP | Transfer from AT5G01380 | Download |
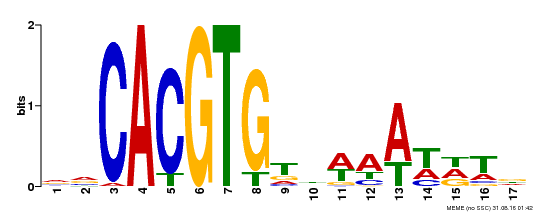
| |||
| Regulation -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | INDUCTION: By salt and infection with the bacterial pathogen P.syringae pv tomato. {ECO:0000269|PubMed:15310827}. | |||||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_017220687.1 | 0.0 | PREDICTED: trihelix transcription factor GT-3b-like | ||||
| Swissprot | O80450 | 5e-69 | TGT3B_ARATH; Trihelix transcription factor GT-3b | ||||
| Swissprot | Q9SDW0 | 6e-69 | TGT3A_ARATH; Trihelix transcription factor GT-3a | ||||
| TrEMBL | A0A175YKF1 | 0.0 | A0A175YKF1_DAUCS; Uncharacterized protein | ||||
| STRING | XP_010264622.1 | 1e-110 | (Nelumbo nucifera) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Asterids | OGEA4187 | 23 | 43 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT5G01380.1 | 7e-51 | Trihelix family protein | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | DCAR_028858 |
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



