 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Cucsa.359570.1 | ||||||||
| Common Name | Csa_7G447800, LOC101215755 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; fabids; Cucurbitales; Cucurbitaceae; Benincaseae; Cucumis
|
||||||||
| Family | GATA | ||||||||
| Protein Properties | Length: 304aa MW: 33950.8 Da PI: 6.8848 | ||||||||
| Description | GATA family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | GATA | 53.3 | 3.8e-17 | 231 | 267 | 1 | 35 |
GATA 1 CsnCgttk..TplWRrgpdgnktLCnaCGlyyrkkgl 35
C++Cg ++ Tp++Rrgpdg++tLCnaCGl+++ kg+
Cucsa.359570.1 231 CRHCGISEksTPMMRRGPDGPRTLCNACGLMWANKGT 267
*****99999************************997 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| PROSITE profile | PS51320 | 15.095 | 78 | 113 | IPR010399 | Tify domain |
| SMART | SM00979 | 3.6E-11 | 78 | 113 | IPR010399 | Tify domain |
| Pfam | PF06200 | 3.7E-13 | 84 | 112 | IPR010399 | Tify domain |
| PROSITE profile | PS51017 | 13.205 | 155 | 197 | IPR010402 | CCT domain |
| Pfam | PF06203 | 6.4E-16 | 155 | 197 | IPR010402 | CCT domain |
| PROSITE profile | PS50114 | 9.249 | 225 | 286 | IPR000679 | Zinc finger, GATA-type |
| SMART | SM00401 | 4.9E-11 | 225 | 278 | IPR000679 | Zinc finger, GATA-type |
| SuperFamily | SSF57716 | 3.94E-13 | 226 | 289 | No hit | No description |
| Gene3D | G3DSA:3.30.50.10 | 1.6E-15 | 230 | 272 | IPR013088 | Zinc finger, NHR/GATA-type |
| Pfam | PF00320 | 8.4E-15 | 231 | 267 | IPR000679 | Zinc finger, GATA-type |
| CDD | cd00202 | 5.55E-15 | 231 | 286 | No hit | No description |
| PROSITE pattern | PS00344 | 0 | 231 | 258 | IPR000679 | Zinc finger, GATA-type |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0006355 | Biological Process | regulation of transcription, DNA-templated | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| GO:0005515 | Molecular Function | protein binding | ||||
| GO:0008270 | Molecular Function | zinc ion binding | ||||
| GO:0043565 | Molecular Function | sequence-specific DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 304 aa Download sequence Send to blast |
MDSIHPSDGR MHIGHAQHSM HTQSVQEQEH HDLHYMSNGN GLADEHENEG HGIMVVEREV 60 QSDHGDLAEN RGVMVDRGGE NCDQLTLSYQ GQVYVFDSVS PEKVQAVLLL LGGREVPLRV 120 PSIPITNQPN DRHLTDEAFN QALANIPPRL SVPQRLASLI RFREKRKERN FDKKIRYTVR 180 KEVALRMQRN KGQFTSSKPI HEDSSLAMAS WEQNESWSSD GNGSQQQEIL CRHCGISEKS 240 TPMMRRGPDG PRTLCNACGL MWANKGTLRD LSKTPNQGGQ TATFNRNENA VQNGDSESNP 300 KGV* |
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Transcriptional activator that specifically binds 5'-GATA-3' or 5'-GAT-3' motifs within gene promoters. {ECO:0000250, ECO:0000269|PubMed:14966217}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00369 | DAP | Transfer from AT3G21175 | Download |
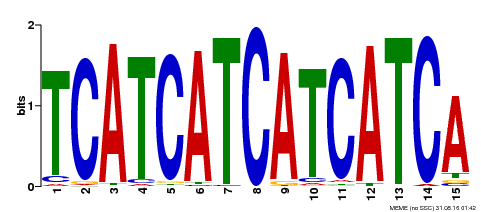
| |||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | HG003874 | 0.0 | HG003874.1 Cucumis sativus mRNA for GATA transcription factor 24-like, cultivar Summer Green. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_011658540.1 | 0.0 | PREDICTED: GATA transcription factor 24-like isoform X2 | ||||
| Swissprot | Q8GXL7 | 1e-117 | GAT24_ARATH; GATA transcription factor 24 | ||||
| TrEMBL | A0A0A0K7I4 | 0.0 | A0A0A0K7I4_CUCSA; Uncharacterized protein | ||||
| STRING | XP_004159006.1 | 0.0 | (Cucumis sativus) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Fabids | OGEF13651 | 23 | 25 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT3G21175.1 | 1e-120 | ZIM-like 1 | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | Cucsa.359570.1 |
| Entrez Gene | 101215755 |
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



