 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Cucsa.288680.1 | ||||||||
| Common Name | Csa_1G575100, LOC101216381 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; fabids; Cucurbitales; Cucurbitaceae; Benincaseae; Cucumis
|
||||||||
| Family | NAC | ||||||||
| Protein Properties | Length: 480aa MW: 53479.3 Da PI: 7.0153 | ||||||||
| Description | NAC family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | NAM | 115.9 | 4e-36 | 48 | 190 | 1 | 128 |
NAM 1 lppGfrFhPtdeelvveyLkkkvegkklelee.......vikev.diykvePwdLpkkvkaeekewyfFskrdkkyatgkrknratk........ 79
lp+G++F+Ptd+el+ e+L++kv++k+++ + +i+ +i++++P++Lp v+ + + +fF++ +k+y+tg+rk+r+++
Cucsa.288680.1 48 LPAGVKFDPTDQELI-EHLEAKVKSKDMKSHPlidefipTIEGEdGICYTHPEKLP-GVTRDGLSRHFFHRPSKAYTTGTRKRRKIQtecdlqgg 140
799************.99*********5553344444434666646**********.555666799******************73334555544 PP
NAM 80 sgyWkatgkdkevlskkgelvglkktLvfy..kgrapkgektdWvmheyrl 128
+ +W++tgk+++v++ +g+++g+kk+Lv+y g+++k+ekt+Wvmh+y+l
Cucsa.288680.1 141 ETRWHKTGKTRPVMA-NGKQKGCKKILVLYtnFGKNRKPEKTNWVMHQYHL 190
778************.999***********5547999999*********98 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| SuperFamily | SSF101941 | 1.7E-38 | 44 | 209 | IPR003441 | NAC domain |
| PROSITE profile | PS51005 | 33.19 | 48 | 209 | IPR003441 | NAC domain |
| Pfam | PF02365 | 2.2E-16 | 49 | 190 | IPR003441 | NAC domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0006355 | Biological Process | regulation of transcription, DNA-templated | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0044212 | Molecular Function | transcription regulatory region DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 480 aa Download sequence Send to blast |
MSGGSGGGSG GGSDSLLDAK LEQHQMCGSK HCPGCGHKLE GRPDWVGLPA GVKFDPTDQE 60 LIEHLEAKVK SKDMKSHPLI DEFIPTIEGE DGICYTHPEK LPGVTRDGLS RHFFHRPSKA 120 YTTGTRKRRK IQTECDLQGG ETRWHKTGKT RPVMANGKQK GCKKILVLYT NFGKNRKPEK 180 TNWVMHQYHL GQHEEEKEGE LVVSKIFYQT QPRQCNWSER SMAAAEGSFD VSNLSRRETS 240 AVTTGSCSSM TQADDVSPAA TTGVGCSISS FSSLDIQHLK SDHFGFVPFR TTFDEVGMEE 300 GSTERKIGSK DGRGGSGEFE LRDHRHQRAA TLDHHHHHHH HQLVVAHDDH HHHSVVNHHL 360 PTTTFHVTNP THQISSIVIS PPPLLLLDHD SYHHHHSPII LQNQPFHQEQ EQQEEGESEH 420 HKMGGRSASG LEELIMGCTS SSIKHQPESS MPSGRETDQW MKYSSFWPDP NNPNLHGHG* |
| Nucleic Localization Signal ? help Back to Top | |||
|---|---|---|---|
| No. | Start | End | Sequence |
| 1 | 1 | 12 | SGGSGGGSGGGS |
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Transcription activator involved in xylem formation (PubMed:25265867, PubMed:25535195). Promotes the expression of the secondary wall-associated transcription factor MYB46 (PubMed:25265867). Functions upstream of NAC030/VND7, a master switch of xylem vessel differentiation (PubMed:25265867, PubMed:25535195). Acts as upstream regulator of NAC101/VND6 and LBD30/ASL19 (PubMed:25535195). {ECO:0000269|PubMed:25265867, ECO:0000269|PubMed:25535195}. | |||||
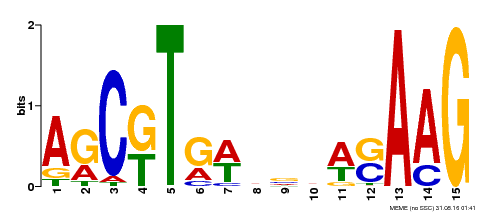
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00460 | DAP | Transfer from AT4G29230 | Download |
| |||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | LN681806 | 0.0 | LN681806.1 Cucumis melo genomic scaffold, anchoredscaffold00025. | |||
| GenBank | LN713256 | 0.0 | LN713256.1 Cucumis melo genomic chromosome, chr_2. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_004135810.2 | 0.0 | PREDICTED: uncharacterized protein LOC101216381 isoform X2 | ||||
| Swissprot | Q9M0F8 | 1e-167 | NAC75_ARATH; NAC domain-containing protein 75 | ||||
| TrEMBL | A0A0A0LYR7 | 0.0 | A0A0A0LYR7_CUCSA; Uncharacterized protein | ||||
| STRING | XP_004158457.1 | 0.0 | (Cucumis sativus) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Fabids | OGEF4520 | 34 | 58 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT4G29230.1 | 1e-138 | NAC domain containing protein 75 | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | Cucsa.288680.1 |
| Entrez Gene | 101216381 |
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



