 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Cucsa.241900.1 | ||||||||
| Common Name | Csa_3G782640, LOC101218755 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; fabids; Cucurbitales; Cucurbitaceae; Benincaseae; Cucumis
|
||||||||
| Family | Dof | ||||||||
| Protein Properties | Length: 212aa MW: 23519.4 Da PI: 7.261 | ||||||||
| Description | Dof family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | zf-Dof | 118.5 | 2.5e-37 | 14 | 66 | 7 | 59 |
zf-Dof 7 kcprCdstntkfCyynnyslsqPryfCkaCrryWtkGGalrnvPvGggrrknk 59
+cprC s+ntkfCyynnysl+qPryfCk+CrryWtkGG+lrnvPvGgg+rkn+
Cucsa.241900.1 14 NCPRCGSSNTKFCYYNNYSLTQPRYFCKGCRRYWTKGGSLRNVPVGGGCRKNR 66
8**************************************************98 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| ProDom | PD007478 | 2.0E-26 | 1 | 66 | IPR003851 | Zinc finger, Dof-type |
| Pfam | PF02701 | 6.1E-32 | 13 | 66 | IPR003851 | Zinc finger, Dof-type |
| PROSITE profile | PS50884 | 28.379 | 13 | 67 | IPR003851 | Zinc finger, Dof-type |
| PROSITE pattern | PS01361 | 0 | 15 | 51 | IPR003851 | Zinc finger, Dof-type |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0006355 | Biological Process | regulation of transcription, DNA-templated | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 212 aa Download sequence Send to blast |
MERGWKPMVE ISPNCPRCGS SNTKFCYYNN YSLTQPRYFC KGCRRYWTKG GSLRNVPVGG 60 GCRKNRLVYA NFVNQKPDTP ETIPTSNAAG HFPVSVQSRV VGCGNFCEFS MQECGLGAND 120 YQVGGGCFGL EESSNNNESQ VAGRELQLPP LPGEDMVEAW GNSMIMVNNH HRLQPTRVEM 180 FEDPNLLSGN WSPIDLSNYD TFSKGLTNIH I* |
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
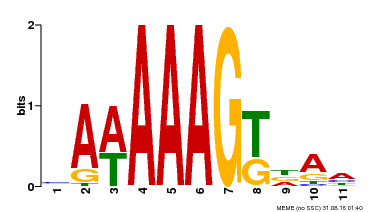
| UniProt | Transcription factor that binds specifically to a 5'-AA[AG]G-3' consensus core sequence. {ECO:0000250}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00405 | DAP | Transfer from AT3G52440 | Download |
| |||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | LN681841 | 1e-156 | LN681841.1 Cucumis melo genomic scaffold, anchoredscaffold00011. | |||
| GenBank | LN713258 | 1e-156 | LN713258.1 Cucumis melo genomic chromosome, chr_4. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_004136713.1 | 1e-149 | PREDICTED: dof zinc finger protein DOF3.5 | ||||
| Swissprot | Q9SVC5 | 3e-43 | DOF35_ARATH; Dof zinc finger protein DOF3.5 | ||||
| TrEMBL | A0A0A0LGT7 | 1e-147 | A0A0A0LGT7_CUCSA; Uncharacterized protein | ||||
| STRING | XP_004136713.1 | 1e-148 | (Cucumis sativus) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Fabids | OGEF1817 | 33 | 94 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT3G52440.1 | 1e-45 | Dof family protein | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | Cucsa.241900.1 |
| Entrez Gene | 101218755 |
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



