 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Cucsa.219780.1 | ||||||||
| Common Name | Csa_7G073570, LOC101203950 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; fabids; Cucurbitales; Cucurbitaceae; Benincaseae; Cucumis
|
||||||||
| Family | bZIP | ||||||||
| Protein Properties | Length: 410aa MW: 44875.6 Da PI: 10.1491 | ||||||||
| Description | bZIP family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | bZIP_1 | 43.3 | 8e-14 | 331 | 381 | 5 | 55 |
CHHHCHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHH CS
bZIP_1 5 krerrkqkNReAArrsRqRKkaeieeLeekvkeLeaeNkaLkkeleelkke 55
+r+rr++kNRe+A rsR+RK+a++ eLe +v++L++ N++L+k+ e+ ++
Cucsa.219780.1 331 RRQRRMIKNRESAARSRARKQAYTLELEAEVAKLKEMNQELQKKQREIMET 381
79****************************************998887765 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| SMART | SM00338 | 2.8E-12 | 327 | 391 | IPR004827 | Basic-leucine zipper domain |
| PROSITE profile | PS50217 | 10.703 | 329 | 380 | IPR004827 | Basic-leucine zipper domain |
| CDD | cd14707 | 1.39E-25 | 331 | 385 | No hit | No description |
| SuperFamily | SSF57959 | 1.34E-9 | 331 | 380 | No hit | No description |
| Pfam | PF00170 | 1.4E-11 | 331 | 383 | IPR004827 | Basic-leucine zipper domain |
| Gene3D | G3DSA:1.20.5.170 | 1.9E-13 | 331 | 382 | No hit | No description |
| PROSITE pattern | PS00036 | 0 | 334 | 349 | IPR004827 | Basic-leucine zipper domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0009414 | Biological Process | response to water deprivation | ||||
| GO:0009651 | Biological Process | response to salt stress | ||||
| GO:0009738 | Biological Process | abscisic acid-activated signaling pathway | ||||
| GO:0045893 | Biological Process | positive regulation of transcription, DNA-templated | ||||
| GO:0000976 | Molecular Function | transcription regulatory region sequence-specific DNA binding | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 410 aa Download sequence Send to blast |
MNFRNFEDIP PGEGPMAKAQ GNFTLTRQPS IYSLTFDEFQ NTWNGLGKDV GSMNMDELLK 60 NIWTAEESQA VTSAGAATGG AGITNGGNLQ RQGSLTLPRT ISQKTVDEVW KDLSKENTSV 120 NEGNGIEPMP ARRQPTLGEV TLEEFLARAG VVREEPPHIE ERPFNCGFYG GLSREDNNGS 180 LALGMFMGNQ ITENKNMVSN QNQNPVFLGT GVVRSSQPQQ QQQPLFPKPA NVTFASSMNL 240 VNNPQLTNGS GTNLVVAPKP PLHDALIQGT GIGAIGLGTR GVTVASRSPT STISSDVITK 300 SSIEASSFSP VPFSFGRGRR SSGALEKVVE RRQRRMIKNR ESAARSRARK QAYTLELEAE 360 VAKLKEMNQE LQKKQREIME TQKNQVLEKM KYQLGGKRFC LRRTLTGPW* |
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Binds to the ABA-responsive element (ABRE). Mediates stress-responsive ABA signaling. {ECO:0000269|PubMed:11884679, ECO:0000269|PubMed:15361142}. | |||||
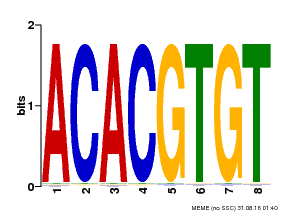
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00038 | PBM | Transfer from AT4G34000 | Download |
| |||
| Regulation -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | INDUCTION: Up-regulated by drought, salt, abscisic acid (ABA). {ECO:0000269|PubMed:10636868, ECO:0000269|PubMed:16284313}. | |||||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | LN681792 | 0.0 | LN681792.1 Cucumis melo genomic scaffold, anchoredscaffold00034. | |||
| GenBank | LN713255 | 0.0 | LN713255.1 Cucumis melo genomic chromosome, chr_1. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_004136974.1 | 0.0 | PREDICTED: ABSCISIC ACID-INSENSITIVE 5-like protein 6 | ||||
| Refseq | XP_011658897.1 | 0.0 | PREDICTED: ABSCISIC ACID-INSENSITIVE 5-like protein 6 | ||||
| Swissprot | Q9M7Q3 | 1e-108 | AI5L6_ARATH; ABSCISIC ACID-INSENSITIVE 5-like protein 6 | ||||
| TrEMBL | A0A0A0K7Y6 | 0.0 | A0A0A0K7Y6_CUCSA; Uncharacterized protein | ||||
| STRING | XP_004136974.1 | 0.0 | (Cucumis sativus) | ||||
| STRING | XP_004159895.1 | 0.0 | (Cucumis sativus) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Fabids | OGEF1962 | 34 | 81 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT4G34000.2 | 5e-97 | abscisic acid responsive elements-binding factor 3 | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | Cucsa.219780.1 |
| Entrez Gene | 101203950 |



