 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Cucsa.118340.1 | ||||||||
| Common Name | Csa_6G139160, LOC101221336 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; fabids; Cucurbitales; Cucurbitaceae; Benincaseae; Cucumis
|
||||||||
| Family | NAC | ||||||||
| Protein Properties | Length: 313aa MW: 35728.8 Da PI: 8.0699 | ||||||||
| Description | NAC family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | NAM | 160.8 | 5.4e-50 | 15 | 142 | 3 | 128 |
NAM 3 pGfrFhPtdeelvveyLkkkvegkkleleevikevdiykvePwdLp.kkvkaeekewyfFskrdkkyatgkrknratksgyWkatgkdkevlsk. 95
pGfrFhPtdeelv +yL++kv++k + + e ik++diyk++PwdLp + + +ekewyfF+kr +ky+++ r+nr+t sg+Wkatg dk++++
Cucsa.118340.1 15 PGFRFHPTDEELVGFYLRRKVDKKAIGT-ELIKSIDIYKHNPWDLPnGSNNLGEKEWYFFCKRGRKYKNSIRPNRVTGSGFWKATGIDKAIYNGs 108
9************************999.89***************445556999*************************************976 PP
NAM 96 .kgelvglkktLvfykgrapkgektdWvmheyrl 128
+++ +glkktLv+ykg+a +g+kt+W+mhe+rl
Cucsa.118340.1 109 qGSNIIGLKKTLVYYKGNAGRGTKTEWMMHEFRL 142
67777***************************98 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| SuperFamily | SSF101941 | 3.27E-53 | 12 | 177 | IPR003441 | NAC domain |
| PROSITE profile | PS51005 | 52.959 | 13 | 177 | IPR003441 | NAC domain |
| Pfam | PF02365 | 2.1E-26 | 15 | 142 | IPR003441 | NAC domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0006355 | Biological Process | regulation of transcription, DNA-templated | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 313 aa Download sequence Send to blast |
MFAEPEIDDY QFPLPGFRFH PTDEELVGFY LRRKVDKKAI GTELIKSIDI YKHNPWDLPN 60 GSNNLGEKEW YFFCKRGRKY KNSIRPNRVT GSGFWKATGI DKAIYNGSQG SNIIGLKKTL 120 VYYKGNAGRG TKTEWMMHEF RLPNPHTTTS SIFGRITTSS NKLQDAEIWT LCRIFKRSVS 180 STKYAPNWRE IAGGNGGSIF GEMNNNNGNA YSEECYDNES NYISFSSSLI NFEEKKPIIN 240 NNNNNHNPLM MMMNFSSISE EQEAPSSSTI EELSPSSYSN FDGDATDLFG YKDKNKLQLL 300 NQDFSSFIHH HR* |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 3swm_A | 4e-48 | 15 | 183 | 22 | 174 | NAC domain-containing protein 19 |
| 3swm_B | 4e-48 | 15 | 183 | 22 | 174 | NAC domain-containing protein 19 |
| 3swm_C | 4e-48 | 15 | 183 | 22 | 174 | NAC domain-containing protein 19 |
| 3swm_D | 4e-48 | 15 | 183 | 22 | 174 | NAC domain-containing protein 19 |
| 3swp_A | 4e-48 | 15 | 183 | 22 | 174 | NAC domain-containing protein 19 |
| 3swp_B | 4e-48 | 15 | 183 | 22 | 174 | NAC domain-containing protein 19 |
| 3swp_C | 4e-48 | 15 | 183 | 22 | 174 | NAC domain-containing protein 19 |
| 3swp_D | 4e-48 | 15 | 183 | 22 | 174 | NAC domain-containing protein 19 |
| Search in ModeBase | ||||||
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Transcription factor that binds to the 5'- RRYGCCGT-3' consensus core sequence. Central longevity regulator. Negative regulator of leaf senescence. Modulates cellular H(2)O(2) levels and enhances tolerance to various abiotic stresses through the regulation of DREB2A. {ECO:0000269|PubMed:22345491}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00353 | DAP | Transfer from AT3G12910 | Download |
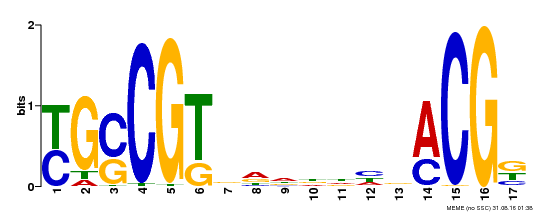
| |||
| Regulation -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | INDUCTION: Up-regulated by H(2)O(2), paraquat, ozone, 3-aminotriazole and salt stress. {ECO:0000269|PubMed:22345491}. | |||||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | LN681921 | 0.0 | LN681921.1 Cucumis melo genomic scaffold, anchoredscaffold00039. | |||
| GenBank | LN713265 | 0.0 | LN713265.1 Cucumis melo genomic chromosome, chr_11. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_011656985.1 | 0.0 | PREDICTED: transcription factor JUNGBRUNNEN 1-like | ||||
| Swissprot | Q9SK55 | 2e-75 | NAC42_ARATH; Transcription factor JUNGBRUNNEN 1 | ||||
| TrEMBL | A0A0A0KGC6 | 0.0 | A0A0A0KGC6_CUCSA; Uncharacterized protein | ||||
| STRING | XP_008456332.1 | 0.0 | (Cucumis melo) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Fabids | OGEF1013 | 34 | 112 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT3G12910.1 | 5e-81 | NAC family protein | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | Cucsa.118340.1 |
| Entrez Gene | 101221336 |



