 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Csa14g043180.1 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; malvids; Brassicales; Brassicaceae; Camelineae; Camelina
|
||||||||
| Family | Trihelix | ||||||||
| Protein Properties | Length: 866aa MW: 95036.8 Da PI: 6.0503 | ||||||||
| Description | Trihelix family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | trihelix | 37.7 | 5.2e-12 | 141 | 181 | 1 | 41 |
trihelix 1 rWtkqevlaLiearremeerlrrgklkkplWeevskkmrer 41
rW+++e+laL+++r++m++++r++ lk+plWe+vs+ re+
Csa14g043180.1 141 RWPREETLALLRIRSDMDSTFRDATLKAPLWEHVSREEREK 181
8***********************************99775 PP
| |||||||
| 2 | trihelix | 96.8 | 2e-30 | 255 | 339 | 1 | 87 |
trihelix 1 rWtkqevlaLiearremeerlrrgklkkplWeevskkmrergferspkqCkekwenlnkrykkikegekkrtsessstcpyfdqlea 87
rW+++e+laL+++r++m++++r++ lk+plWe+vs+k+ e g++rs+k+Ckek+en++k+yk++ke++++r++++ +++f+qlea
Csa14g043180.1 255 RWPREETLALLRIRSDMDSTFRDATLKAPLWEHVSRKLLELGYKRSAKKCKEKFENVQKYYKRTKETRGGRHDGK--AYKFFSQLEA 339
8*********************************************************************86555..5******985 PP
| |||||||
| 3 | trihelix | 103.8 | 1.3e-32 | 629 | 713 | 1 | 86 |
trihelix 1 rWtkqevlaLiearremeerlrrgklkkplWeevskkmrergferspkqCkekwenlnkrykkikegekkrtsessstcpyfdqle 86
rW+k e+laLi++r+ me r++++ k+ lWee+s m++ g++r++k+Ckekwen+nk+ykk+ke++kkr +++ +tcpyf++l+
Csa14g043180.1 629 RWPKAEILALINLRSGMEPRYQDNVPKGLLWEEISTSMKRMGYNRNAKRCKEKWENINKYYKKVKESNKKR-PQDAKTCPYFHRLD 713
8*********************************************************************8.99999*******97 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| PROSITE profile | PS50090 | 4.054 | 134 | 195 | IPR017877 | Myb-like domain |
| PROSITE profile | PS50090 | 6.864 | 248 | 312 | IPR017877 | Myb-like domain |
| SMART | SM00717 | 0.0076 | 252 | 314 | IPR001005 | SANT/Myb domain |
| CDD | cd12203 | 9.21E-26 | 254 | 319 | No hit | No description |
| Pfam | PF13837 | 1.5E-20 | 254 | 340 | No hit | No description |
| SMART | SM00717 | 0.011 | 626 | 688 | IPR001005 | SANT/Myb domain |
| Pfam | PF13837 | 8.3E-22 | 628 | 713 | No hit | No description |
| PROSITE profile | PS50090 | 6.667 | 628 | 686 | IPR017877 | Myb-like domain |
| CDD | cd12203 | 1.56E-28 | 629 | 693 | No hit | No description |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0008361 | Biological Process | regulation of cell size | ||||
| GO:0010090 | Biological Process | trichome morphogenesis | ||||
| GO:0030308 | Biological Process | negative regulation of cell growth | ||||
| GO:0032876 | Biological Process | negative regulation of DNA endoreduplication | ||||
| GO:0042631 | Biological Process | cellular response to water deprivation | ||||
| GO:0045892 | Biological Process | negative regulation of transcription, DNA-templated | ||||
| GO:2000037 | Biological Process | regulation of stomatal complex patterning | ||||
| GO:2000038 | Biological Process | regulation of stomatal complex development | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0043565 | Molecular Function | sequence-specific DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 866 aa Download sequence Send to blast |
MEQVGGGGGG NEVVEEASPI SSRPPANLEE LMRFSAAAAD DGGGGGGGGG KILNQTVRAL 60 AREEREKEKR LVRPSRLVKS RDMEQVGGGG GGNEVVEEAS PISSRPPANL EELMRFSAAA 120 ADDGGGGGGG GGSASSSSGN RWPREETLAL LRIRSDMDST FRDATLKAPL WEHVSREERE 180 KEKRLVRPSR LVKSRDMEQV GGGGGGNEVV EEASPISSRP PANLEELMRF SAAAADDGGG 240 GGGGGGSASS SSGNRWPREE TLALLRIRSD MDSTFRDATL KAPLWEHVSR KLLELGYKRS 300 AKKCKEKFEN VQKYYKRTKE TRGGRHDGKA YKFFSQLEAL NTTPTPPPSH PPSSSLDVTP 360 LSVANPILMP TSSSSPFPVF SQPQTQPPQT HTVTFTPTPQ PPPMAPTFPG VAFSSHSSST 420 ASGMGSDDDE EDDMGVDQAN IAGSSSRKRK RGNHGGGKMM ELFEGLVRQV MQKQAAMQRS 480 FLEALEKREQ ERLDREEAWK RQQMSRLARE HEVMSQERAA SASRDAAIIS LIQKITGHTI 540 QLPPSLSSQT PQPPHQPPQP PPAAKRAQEP PLSTAQSQLQ QPIMAIPQQQ ILPHLPHQPE 600 QKQQQQQQQQ QQQEMMVSSE QSSLPSSSRW PKAEILALIN LRSGMEPRYQ DNVPKGLLWE 660 EISTSMKRMG YNRNAKRCKE KWENINKYYK KVKESNKKRP QDAKTCPYFH RLDLLYRNKV 720 LGGGGGTSTS SGLPQEQKQS PVSAVKPPQE GVVNVQPHGS AASSEELEPI EESPQGTEKP 780 EDLVMRELMQ QQQQQQQQQQ ESMIGEYEKI EESHNYNNME EEEEEMDEEE LDEDEKSAAF 840 EIAFQSPANR GGNGHTEPPF LTMVQ* |
| Nucleic Localization Signal ? help Back to Top | |||
|---|---|---|---|
| No. | Start | End | Sequence |
| 1 | 297 | 305 | KRSAKKCKE |
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
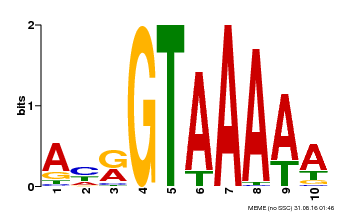
| UniProt | Transcription repressor that binds specific DNA sequence such as GT3 box 5'-GGTAAA-3' in the SDD1 promoter. Negative regulator of water use efficiency (WUE) via the promotion of stomatal density and distribution by the transcription repression of SDD1. Regulates the expression of several cell cycle genes and endoreduplication, especially in trichomes where it prevents ploidy-dependent plant cell growth. {ECO:0000269|PubMed:19717615, ECO:0000269|PubMed:21169508}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00177 | DAP | Transfer from AT1G33240 | Download |
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | Csa14g043180.1 |
| Regulation -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | INDUCTION: Down-regulated by water stress. {ECO:0000269|PubMed:21169508}. | |||||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | AC021045 | 0.0 | AC021045.2 Arabidopsis thaliana chromosome I BAC T9L6 genomic sequence, complete sequence. | |||
| GenBank | AC027035 | 0.0 | AC027035.5 Arabidopsis thaliana chromosome 1 BAC T16O9 genomic sequence, complete sequence. | |||
| GenBank | AJ003215 | 0.0 | AJ003215.1 Arabidopsis thaliana GTL1 gene. | |||
| GenBank | CP002684 | 0.0 | CP002684.1 Arabidopsis thaliana chromosome 1 sequence. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_010461138.1 | 0.0 | PREDICTED: trihelix transcription factor GTL1-like isoform X1 | ||||
| Refseq | XP_019091258.1 | 0.0 | PREDICTED: trihelix transcription factor GTL1-like isoform X2 | ||||
| Swissprot | Q9C882 | 0.0 | GTL1_ARATH; Trihelix transcription factor GTL1 | ||||
| TrEMBL | A0A397Y2Y6 | 0.0 | A0A397Y2Y6_BRACM; Uncharacterized protein | ||||
| TrEMBL | M4FFY6 | 0.0 | M4FFY6_BRARP; Uncharacterized protein | ||||
| STRING | XP_010461140.1 | 0.0 | (Camelina sativa) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Malvids | OGEM6226 | 27 | 46 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT1G33240.1 | 4e-97 | GT-2-like 1 | ||||
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



