 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Cotton_A_37867_BGI-A2_v1.0 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; malvids; Malvales; Malvaceae; Malvoideae; Gossypium
|
||||||||
| Family | C2H2 | ||||||||
| Protein Properties | Length: 565aa MW: 61416.3 Da PI: 8.9594 | ||||||||
| Description | C2H2 family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | zf-C2H2 | 16.2 | 2.9e-05 | 65 | 87 | 1 | 23 |
EEETTTTEEESSHHHHHHHHHHT CS
zf-C2H2 1 ykCpdCgksFsrksnLkrHirtH 23
+ C+ C+k F r nL+ H r H
Cotton_A_37867_BGI-A2_v1.0 65 FICEVCNKGFQREQNLQLHRRGH 87
89*******************88 PP
| |||||||
| 2 | zf-C2H2 | 13.2 | 0.00027 | 141 | 163 | 1 | 23 |
EEETTTTEEESSHHHHHHHHHHT CS
zf-C2H2 1 ykCpdCgksFsrksnLkrHirtH 23
+kC++C+k++ +s+ k H +t+
Cotton_A_37867_BGI-A2_v1.0 141 WKCEKCSKRYAVQSDWKAHSKTC 163
58*****************9998 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| Gene3D | G3DSA:3.30.160.60 | 6.3E-6 | 64 | 87 | IPR013087 | Zinc finger C2H2-type/integrase DNA-binding domain |
| SuperFamily | SSF57667 | 1.67E-7 | 64 | 87 | No hit | No description |
| SMART | SM00355 | 0.0052 | 65 | 87 | IPR015880 | Zinc finger, C2H2-like |
| PROSITE profile | PS50157 | 10.99 | 65 | 87 | IPR007087 | Zinc finger, C2H2 |
| Pfam | PF12171 | 2.8E-5 | 65 | 87 | IPR022755 | Zinc finger, double-stranded RNA binding |
| PROSITE pattern | PS00028 | 0 | 67 | 87 | IPR007087 | Zinc finger, C2H2 |
| SMART | SM00355 | 130 | 106 | 136 | IPR015880 | Zinc finger, C2H2-like |
| Gene3D | G3DSA:3.30.160.60 | 2.0E-5 | 129 | 162 | IPR013087 | Zinc finger C2H2-type/integrase DNA-binding domain |
| SuperFamily | SSF57667 | 1.67E-7 | 136 | 161 | No hit | No description |
| SMART | SM00355 | 130 | 141 | 161 | IPR015880 | Zinc finger, C2H2-like |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0003676 | Molecular Function | nucleic acid binding | ||||
| GO:0046872 | Molecular Function | metal ion binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 565 aa Download sequence Send to blast |
MAASSSLGPF GIREEDQHQH STAAPTSAMG PAPPPPQRKK RNQPVHPDVE VIALSPKSLM 60 ATNRFICEVC NKGFQREQNL QLHRRGHNLP WKLRQKTTKE VKRKVYLCPE PTCVHHDPSR 120 ALGDLTGIKK HYSRKHGEKK WKCEKCSKRY AVQSDWKAHS KTCGTREYRC DCGTLFSRRD 180 SFITHRAFCD ALAQENARQP PSFNSIGNHL YGSTGNICLG LSQVGTQMSS VQDQANNNSG 240 GRDIFKLGGV ARSTQFDHLL LSPSMGSSSS SSFRPQQSIV SSAAFFMPES NQEHHPQQGI 300 LGNNKQYHQG LMMQFPADIQ NNTTNTPSPP SFFNLSFLSN CGNDINLDAN LTSSGLLTPE 360 HFKNETGGGG GGTSEASDPF SNTVMGNQIT TNIPSVFAQS NNIAVQMSAT ALLQKAAQMG 420 STSSYKNASL MRSFGNSKFG GTLGDSTGNN LHELMNSIAG GGSNTYSFGH GQENPYANRS 480 SVEQEKHLEQ QPQILNVSGG CGSDRLTRDF LGVGQIVRSM SMGGVSQREQ QQQQEAMGLS 540 ALGSERSNIT APTNHQCLGG NGNFQ |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 5b3h_C | 2e-34 | 137 | 198 | 3 | 64 | Zinc finger protein JACKDAW |
| 5b3h_F | 2e-34 | 137 | 198 | 3 | 64 | Zinc finger protein JACKDAW |
| Search in ModeBase | ||||||
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Transcription factor acting as a positive regulator of the starch synthase SS4. Controls chloroplast development and starch granule formation (PubMed:22898356). Binds DNA via its zinc fingers (PubMed:24821766). Recognizes and binds to SCL3 promoter sequence 5'-AGACAA-3' to promotes its expression when in complex with RGA (PubMed:24821766). {ECO:0000269|PubMed:22898356, ECO:0000269|PubMed:24821766}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
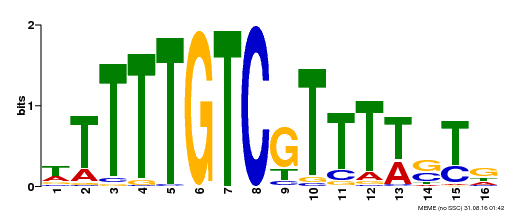
| Motif ID | Method | Source | Motif file |
| MP00255 | DAP | Transfer from AT2G02070 | Download |
| |||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | JX614588 | 0.0 | JX614588.1 Gossypium hirsutum clone NBRI_GE58088 microsatellite sequence. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_017642443.1 | 0.0 | PREDICTED: protein indeterminate-domain 5, chloroplastic-like isoform X1 | ||||
| Swissprot | Q9ZUL3 | 1e-129 | IDD5_ARATH; Protein indeterminate-domain 5, chloroplastic | ||||
| TrEMBL | A0A1U8L014 | 0.0 | A0A1U8L014_GOSHI; protein indeterminate-domain 5, chloroplastic-like isoform X1 | ||||
| STRING | Gorai.002G169100.1 | 0.0 | (Gossypium raimondii) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Malvids | OGEM4373 | 26 | 54 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT2G02070.1 | 1e-113 | indeterminate(ID)-domain 5 | ||||
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



