 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Cotton_A_02413_BGI-A2_v1.0 | ||||||||
| Common Name | F383_23187 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; malvids; Malvales; Malvaceae; Malvoideae; Gossypium
|
||||||||
| Family | MYB_related | ||||||||
| Protein Properties | Length: 299aa MW: 33022.1 Da PI: 9.4607 | ||||||||
| Description | MYB_related family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | Myb_DNA-binding | 40.4 | 6.6e-13 | 39 | 83 | 1 | 47 |
TSSS-HHHHHHHHHHHHHTTTT-HHHHHHHHTTTS-HHHHHHHHHHH CS
Myb_DNA-binding 1 rgrWTteEdellvdavkqlGggtWktIartmgkgRtlkqcksrwqky 47
r +WT+ E++++++a +++ + Wk+I +++g +t q++s+ qky
Cotton_A_02413_BGI-A2_v1.0 39 RESWTEPEHDKFLEALQLFDRD-WKKIEAYVG-SKTVIQIRSHAQKY 83
789*****************77.*********.*************9 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| SuperFamily | SSF46689 | 3.14E-16 | 33 | 89 | IPR009057 | Homeodomain-like |
| PROSITE profile | PS51294 | 14.746 | 34 | 88 | IPR017930 | Myb domain |
| TIGRFAMs | TIGR01557 | 1.0E-18 | 37 | 86 | IPR006447 | Myb domain, plants |
| Gene3D | G3DSA:1.10.10.60 | 6.4E-6 | 38 | 79 | IPR009057 | Homeodomain-like |
| SMART | SM00717 | 2.7E-10 | 38 | 86 | IPR001005 | SANT/Myb domain |
| Pfam | PF00249 | 1.1E-10 | 39 | 83 | IPR001005 | SANT/Myb domain |
| CDD | cd00167 | 6.60E-8 | 41 | 84 | No hit | No description |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0006355 | Biological Process | regulation of transcription, DNA-templated | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 299 aa Download sequence Send to blast |
MALPGLGPFS TAETAASTVS SSEDPNKKIR KPYTISKSRE SWTEPEHDKF LEALQLFDRD 60 WKKIEAYVGS KTVIQIRSHA QKYFLKVQKS GTSEHLPPPR PKRKAAHPYP QKASKNVLGQ 120 PQVSEPLQSP AALLDTGYVL RSNPSLMLAD PVTRAAASPQ TNNAQTISFA QDKKGPGMAN 180 NSCSSTESTR KTTEVGDTTD RGNHGHGLRV LPDFAQVYSF IGSVFDPNTK GHMQKLKKMD 240 PIDVETVLLL MRNLSINLTS PDFEDHRRLL SSYEIDTETN NINSGACEAV DNNQSKRIT |
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Probable transcription factor. {ECO:0000250}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00625 | PBM | Transfer from PK02532.1 | Download |
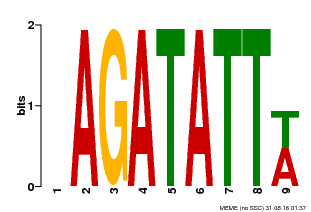
| |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_017608742.1 | 0.0 | PREDICTED: protein REVEILLE 6-like isoform X2 | ||||
| Swissprot | Q8H0W3 | 1e-126 | RVE6_ARATH; Protein REVEILLE 6 | ||||
| TrEMBL | A0A0B0MF46 | 0.0 | A0A0B0MF46_GOSAR; Transcription factor ASG4-like protein | ||||
| STRING | Gorai.007G013200.1 | 0.0 | (Gossypium raimondii) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Malvids | OGEM1290 | 28 | 93 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT5G52660.1 | 1e-111 | MYB_related family protein | ||||
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



