 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Cla023247 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; fabids; Cucurbitales; Cucurbitaceae; Benincaseae; Citrullus
|
||||||||
| Family | bHLH | ||||||||
| Protein Properties | Length: 575aa MW: 62911.8 Da PI: 7.0611 | ||||||||
| Description | bHLH family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | HLH | 51.7 | 1.5e-16 | 356 | 402 | 4 | 55 |
HHHHHHHHHHHHHHHHHHHHHCTSCCC...TTS-STCHHHHHHHHHHHHHHH CS
HLH 4 ahnerErrRRdriNsafeeLrellPkaskapskKlsKaeiLekAveYIksLq 55
hn ErrRR+riN+++ L+el+P++ K +Ka++L +A+eY+ksLq
Cla023247 356 VHNLSERRRRERINEKMKALQELIPHC-----NKTDKASMLDEAIEYLKSLQ 402
6*************************8.....6******************9 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| SuperFamily | SSF47459 | 7.85E-21 | 349 | 416 | IPR011598 | Myc-type, basic helix-loop-helix (bHLH) domain |
| PROSITE profile | PS50888 | 18.59 | 352 | 401 | IPR011598 | Myc-type, basic helix-loop-helix (bHLH) domain |
| Gene3D | G3DSA:4.10.280.10 | 1.5E-20 | 356 | 410 | IPR011598 | Myc-type, basic helix-loop-helix (bHLH) domain |
| Pfam | PF00010 | 6.2E-14 | 356 | 402 | IPR011598 | Myc-type, basic helix-loop-helix (bHLH) domain |
| SMART | SM00353 | 1.2E-18 | 358 | 407 | IPR011598 | Myc-type, basic helix-loop-helix (bHLH) domain |
| CDD | cd00083 | 5.44E-9 | 366 | 406 | No hit | No description |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0009704 | Biological Process | de-etiolation | ||||
| GO:0010161 | Biological Process | red light signaling pathway | ||||
| GO:0010244 | Biological Process | response to low fluence blue light stimulus by blue low-fluence system | ||||
| GO:0010600 | Biological Process | regulation of auxin biosynthetic process | ||||
| GO:0010928 | Biological Process | regulation of auxin mediated signaling pathway | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| GO:0046983 | Molecular Function | protein dimerization activity | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 575 aa Download sequence Send to blast |
MNYCFPDWNF EGDLPPANHK KSMGPEHDDL VELLWRNGQV VLQSQKAKKP SLIANEVRQF 60 QKQNQPLYRS NVSCGNSSNL IQDDETVSWI QDPLDDSFEK EFCSNFFSEL PPADPIEIVK 120 QPIKHFQDDK QTRFGAFDTA THVTFGNSLK KNSKPPAALA EIPMNTMPPP RFQFPDSTRQ 180 AKDLGDLGKL VNFSQVPVPL KGDLGSSNGG RECGNLIQGE GRDCSAMTVG SSHCGSNQVP 240 NPNDLDVSRV STSGFGNAGL SAGLSKEDNR KMVAQGERDK IETMDPTATS SSGGSGSSMD 300 RSRTIGQSTG GNSNKRKGRD GEESECQSET AELESAEGNK AAPRSGSSRR TRAAEVHNLS 360 ERRRRERINE KMKALQELIP HCNKTDKASM LDEAIEYLKS LQLQLQVMWM GSGMAPMMFP 420 GIQHYMSRMA MGMGMAQPSM PSIHNSMQLP RVPIVDQSVS VAPTPNQPMM CQPQIFNPMN 480 YQNQMQNPAL QEQYARLMGF HHMQPTSQPI NVFRFCPSAV LQSQTAAAPG PACGSTAGGS 540 IANDIVNANL VIQLLSKSNI FTKVPVSGGR RGGDE |
| Nucleic Localization Signal ? help Back to Top | |||
|---|---|---|---|
| No. | Start | End | Sequence |
| 1 | 360 | 365 | ERRRRE |
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00606 | ChIP-seq | Transfer from AT2G43010 | Download |
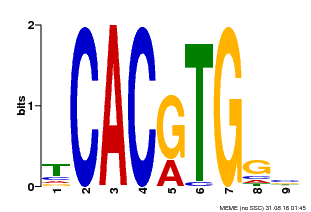
| |||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | LN681864 | 0.0 | LN681864.1 Cucumis melo genomic scaffold, anchoredscaffold00092. | |||
| GenBank | LN713261 | 0.0 | LN713261.1 Cucumis melo genomic chromosome, chr_7. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_011653988.1 | 0.0 | PREDICTED: transcription factor PIF5-like | ||||
| Refseq | XP_011653989.1 | 0.0 | PREDICTED: transcription factor PIF5-like | ||||
| TrEMBL | A0A0A0KZR2 | 0.0 | A0A0A0KZR2_CUCSA; Uncharacterized protein | ||||
| STRING | XP_004162571.1 | 0.0 | (Cucumis sativus) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Fabids | OGEF8146 | 29 | 44 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT2G43010.2 | 5e-61 | phytochrome interacting factor 4 | ||||



