 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Cla015525 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; fabids; Cucurbitales; Cucurbitaceae; Benincaseae; Citrullus
|
||||||||
| Family | Trihelix | ||||||||
| Protein Properties | Length: 375aa MW: 41197.9 Da PI: 10.2731 | ||||||||
| Description | Trihelix family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | trihelix | 50.1 | 7.1e-16 | 70 | 157 | 1 | 86 |
trihelix 1 rWtkqevlaLiearremeerlrrgklkkplWeevskkm....rergferspkqCkekwenlnkrykkikegekkrtsessstcpyfdqle 86
+W++ + +Li+a e+ +l+rg+lk+++W+ev++ + +++ r+ +qCk++++ ++k+yk +k+ + + s++p++d+l+
Cla015525 70 CWSEGATSVLIDAWGERYLELSRGNLKQKHWKEVADIVssreDYTKIPRTDIQCKNRIDTVKKKYKTEKAKITAG--GGPSKWPFYDRLD 157
7*************************************87666677899**********************9997..67779******98 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| Pfam | PF13837 | 2.0E-27 | 68 | 159 | No hit | No description |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0009793 | Biological Process | embryo development ending in seed dormancy | ||||
| GO:0010431 | Biological Process | seed maturation | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 375 aa Download sequence Send to blast |
MKEDDEIQSY PSPGSGSPAS PISNGRITVT VAAAPPPPPS SQNMITLALP NQQSKGGGGG 60 GSGGGGREDC WSEGATSVLI DAWGERYLEL SRGNLKQKHW KEVADIVSSR EDYTKIPRTD 120 IQCKNRIDTV KKKYKTEKAK ITAGGGPSKW PFYDRLDQLI GPASKNSASS AGVATAVNPP 180 LQQNQKVPLG IPVVNRSLIP YQAHHNHSHQ QQPKGPKAQK IQYHKRPRTT DSDSSGSDRE 240 TSPTSSDSYR QGNFQRKNVR VQKEVVNPNL GQVGKTEKGR NGSREKGWKN AVSELTQAIL 300 KFGEAYEQAE SSKLQQVVEM EKQRMKFAKD LELQRMQFFM KTQLEISQLK HGRRVVAAGN 360 HHCSNNNNSN SDSSN |
| Nucleic Localization Signal ? help Back to Top | |||
|---|---|---|---|
| No. | Start | End | Sequence |
| 1 | 54 | 66 | KGGGGGGSGGGGR |
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Transcription regulator that may repress the maturation program during early embryogenesis. {ECO:0000269|PubMed:21330492}. | |||||
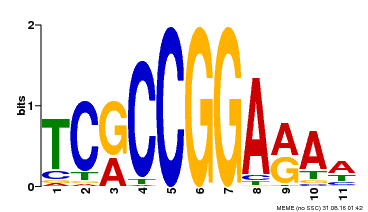
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00356 | DAP | Transfer from AT3G14180 | Download |
| |||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | LN681805 | 0.0 | LN681805.1 Cucumis melo genomic scaffold, anchoredscaffold00067. | |||
| GenBank | LN713255 | 0.0 | LN713255.1 Cucumis melo genomic chromosome, chr_1. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_008462265.1 | 0.0 | PREDICTED: trihelix transcription factor ASIL2 | ||||
| Swissprot | Q9LJG8 | 1e-101 | ASIL2_ARATH; Trihelix transcription factor ASIL2 | ||||
| TrEMBL | A0A1S3CH31 | 0.0 | A0A1S3CH31_CUCME; trihelix transcription factor ASIL2 | ||||
| STRING | XP_008462265.1 | 0.0 | (Cucumis melo) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Fabids | OGEF7919 | 30 | 44 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT3G14180.1 | 8e-80 | sequence-specific DNA binding transcription factors | ||||



