 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Cla009965 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; fabids; Cucurbitales; Cucurbitaceae; Benincaseae; Citrullus
|
||||||||
| Family | bHLH | ||||||||
| Protein Properties | Length: 399aa MW: 43152.2 Da PI: 5.8883 | ||||||||
| Description | bHLH family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | HLH | 31.2 | 3.8e-10 | 233 | 281 | 3 | 55 |
HHHHHHHHHHHHHHHHHHHHHHCTSCCC...TTS-STCHHHHHHHHHHHHHHH CS
HLH 3 rahnerErrRRdriNsafeeLrellPkaskapskKlsKaeiLekAveYIksLq 55
+h+ +Er RR++i +++ L+el+P + +k Ka +L + ++Y++sLq
Cla009965 233 NSHSLAERVRREKISERMRLLQELVPGC----NKITGKAVMLDEIINYVQSLQ 281
58**************************....999*****************9 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| CDD | cd00083 | 2.98E-11 | 228 | 285 | No hit | No description |
| SuperFamily | SSF47459 | 9.95E-17 | 228 | 295 | IPR011598 | Myc-type, basic helix-loop-helix (bHLH) domain |
| PROSITE profile | PS50888 | 15.485 | 230 | 280 | IPR011598 | Myc-type, basic helix-loop-helix (bHLH) domain |
| Gene3D | G3DSA:4.10.280.10 | 1.5E-16 | 231 | 297 | IPR011598 | Myc-type, basic helix-loop-helix (bHLH) domain |
| Pfam | PF00010 | 3.4E-7 | 233 | 281 | IPR011598 | Myc-type, basic helix-loop-helix (bHLH) domain |
| SMART | SM00353 | 4.6E-10 | 236 | 286 | IPR011598 | Myc-type, basic helix-loop-helix (bHLH) domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0009637 | Biological Process | response to blue light | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0046983 | Molecular Function | protein dimerization activity | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 399 aa Download sequence Send to blast |
MGDLENDEMG FRNSNEGLMN CPSTMAMGSF YGSGWDPLVS LGQSENFGAA SSMVSSHGEF 60 SNSFPVVLEN HGGSGTAQQG VQYPTDLEMS TKLPCFGSGN YSDMFGSSFG LAGQILGGCS 120 SNYRNAQTSQ ENLPLSGEET AVGSPNGKIR KRALDSTFPF SPNKKSEVKL KKDVSGDSSS 180 TQEEKNAEME QNLGGNSRGK ATGKQTKEKS NNSAEAPKEN YIHVRARRGQ ATNSHSLAER 240 VRREKISERM RLLQELVPGC NKITGKAVML DEIINYVQSL QQQVEFLSMK LATVNPDVNI 300 DIERILSKDI FHARGSNGTV LGFDPGLSAV SPVPPHRMFQ FQGTMSNMAT TSTQFPPLPQ 360 TMLESDIQSL LQMGFDSGSA LDNLGPNGIV LHNLLALHY |
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00135 | DAP | Transfer from AT1G10120 | Download |
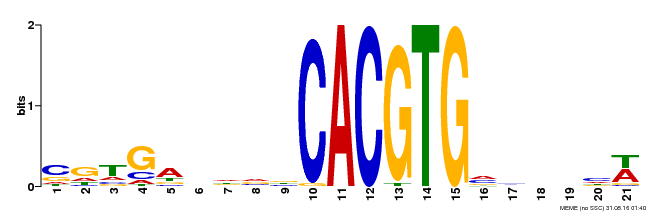
| |||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | LN681861 | 1e-149 | LN681861.1 Cucumis melo genomic scaffold, anchoredscaffold00029. | |||
| GenBank | LN713261 | 1e-149 | LN713261.1 Cucumis melo genomic chromosome, chr_7. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_008452681.1 | 0.0 | PREDICTED: transcription factor bHLH74-like | ||||
| TrEMBL | A0A1S3BV77 | 0.0 | A0A1S3BV77_CUCME; transcription factor bHLH74-like | ||||
| STRING | XP_008452681.1 | 0.0 | (Cucumis melo) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Fabids | OGEF7585 | 33 | 45 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT1G10120.1 | 5e-83 | bHLH family protein | ||||



