 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Ciclev10024892m | ||||||||
| Common Name | CICLE_v10024891mg | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; malvids; Sapindales; Rutaceae; Aurantioideae; Citrus
|
||||||||
| Family | HD-ZIP | ||||||||
| Protein Properties | Length: 839aa MW: 92446.1 Da PI: 6.5047 | ||||||||
| Description | HD-ZIP family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | Homeobox | 58.4 | 1.2e-18 | 17 | 75 | 3 | 57 |
--SS--HHHHHHHHHHHHHSSS--HHHHHHHHHHC....TS-HHHHHHHHHHHHHHHHC CS
Homeobox 3 kRttftkeqleeLeelFeknrypsaeereeLAkkl....gLterqVkvWFqNrRakekk 57
k ++t+eq+e+Le+l++++++ps +r++L +++ +++ +q+kvWFqNrR +ek+
Ciclev10024892m 17 KYVRYTPEQVEALERLYHECPKPSSIRRQQLIRECpilsNIEPKQIKVWFQNRRCREKQ 75
5679*****************************************************97 PP
| |||||||
| 2 | START | 178.6 | 3.8e-56 | 161 | 369 | 2 | 205 |
HHHHHHHHHHHHHHHC-TT-EEEEEXCCTTEEEEEEESSS.SCEEEEEEEECCSCHHHHHHHHHCCCGGCT-TT-SEEEEEEEECTT..EEEEE CS
START 2 laeeaaqelvkkalaeepgWvkssesengdevlqkfeeskvdsgealrasgvvdmvlallveellddkeqWdetlakaetlevissg..galql 93
+aee+++e+++ka+ ++ Wv+++ +++g++++ +++ s+++sg a+ra+g+v +++ v+e+l+d++ W ++++++e+l+v+ ++ g+++l
Ciclev10024892m 161 IAEETLTEFLSKATGTAVEWVQMPGMKPGPDSVGIVAISHGCSGVAARACGLVGLEPT-RVAEILKDRPSWFRDCRAVEVLNVLPTAngGTIEL 253
789*******************************************************.8888888888****************9999***** PP
EEEXXTTXX-SSX.EEEEEEEEEEE.TTS-EEEEEEEEE-TTS--....-TTSEE-EESSEEEEEEEECTCEEEEEEEE-EE--SSXXHHHHHH CS
START 94 mvaelqalsplvp.RdfvfvRyirqlgagdwvivdvSvdseqkppe...sssvvRaellpSgiliepksnghskvtwvehvdlkgrlphwllrs 183
+++l+al++l+p Rdf+ +Ry+ l++g++v++++S+ + q+ p+ +++vRae+lpSg+li+p+++g+s +++v+h+dl+ ++++++lr+
Ciclev10024892m 254 LYMQLYALTTLAPaRDFWLLRYTSVLEDGSLVVCERSLKNIQNGPTmppVQHFVRAEMLPSGYLIRPCEGGGSIIHIVDHMDLEPWSVPEVLRP 347
****************************************99999988899******************************************* PP
HHHHHHHHHHHHHHHHTXXXXX CS
START 184 lvksglaegaktwvatlqrqce 205
l++s+++ ++kt++a+l+++++
Ciclev10024892m 348 LYESSTVLAQKTTMAALRQLRQ 369
*****************99875 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| PROSITE profile | PS50071 | 15.613 | 12 | 76 | IPR001356 | Homeobox domain |
| SMART | SM00389 | 1.2E-15 | 14 | 80 | IPR001356 | Homeobox domain |
| SuperFamily | SSF46689 | 3.64E-17 | 16 | 80 | IPR009057 | Homeodomain-like |
| CDD | cd00086 | 9.31E-17 | 17 | 77 | No hit | No description |
| Pfam | PF00046 | 2.9E-16 | 18 | 75 | IPR001356 | Homeobox domain |
| Gene3D | G3DSA:1.10.10.60 | 1.6E-18 | 19 | 75 | IPR009057 | Homeodomain-like |
| CDD | cd14686 | 1.14E-6 | 69 | 108 | No hit | No description |
| PROSITE profile | PS50848 | 26.049 | 151 | 379 | IPR002913 | START domain |
| CDD | cd08875 | 3.97E-78 | 155 | 369 | No hit | No description |
| Gene3D | G3DSA:3.30.530.20 | 2.4E-24 | 160 | 367 | IPR023393 | START-like domain |
| SuperFamily | SSF55961 | 7.0E-38 | 160 | 372 | No hit | No description |
| SMART | SM00234 | 3.8E-42 | 160 | 370 | IPR002913 | START domain |
| Pfam | PF01852 | 2.2E-53 | 161 | 369 | IPR002913 | START domain |
| SuperFamily | SSF55961 | 4.07E-5 | 397 | 489 | No hit | No description |
| SuperFamily | SSF55961 | 4.07E-5 | 521 | 596 | No hit | No description |
| Pfam | PF08670 | 7.2E-52 | 695 | 837 | IPR013978 | MEKHLA |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0009965 | Biological Process | leaf morphogenesis | ||||
| GO:0010014 | Biological Process | meristem initiation | ||||
| GO:0010075 | Biological Process | regulation of meristem growth | ||||
| GO:0010087 | Biological Process | phloem or xylem histogenesis | ||||
| GO:0048263 | Biological Process | determination of dorsal identity | ||||
| GO:0080060 | Biological Process | integument development | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| GO:0008289 | Molecular Function | lipid binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 839 aa Download sequence Send to blast |
MAMSCKDGKT GSLDNGKYVR YTPEQVEALE RLYHECPKPS SIRRQQLIRE CPILSNIEPK 60 QIKVWFQNRR CREKQRKEAS RLQAVNRKLT AMNKLLMEEN DRLQKQVSQL VYENGYFRQH 120 TQSTTLATKD TSCESVVTSG QHHLTPQHPP RDASPAGLLS IAEETLTEFL SKATGTAVEW 180 VQMPGMKPGP DSVGIVAISH GCSGVAARAC GLVGLEPTRV AEILKDRPSW FRDCRAVEVL 240 NVLPTANGGT IELLYMQLYA LTTLAPARDF WLLRYTSVLE DGSLVVCERS LKNIQNGPTM 300 PPVQHFVRAE MLPSGYLIRP CEGGGSIIHI VDHMDLEPWS VPEVLRPLYE SSTVLAQKTT 360 MAALRQLRQM AQEVTQSSVN GWGRRPAALR ALSQRLSRGF NEAVNGFTDE GWTVMGNDGM 420 DDVTVLVNSS PDKLMGLNLS FANGFPAVSN AVLCAKASML LQNVPPAILL RFLREHRSEW 480 ADNNIDVYSA AAIKVGPCSL PGSRVGTFGS QVILPLAHTI EHEEFMEVIK LEGVGHSPED 540 AIMPRDMFLL QLCSGMDENA VGTCAELIFA PIDASFADDA PLLPSGFRII PLDSGKETSS 600 PNRTLDLASA LEIGPAGNRA TNNYSTNSTC MRSVMTIAFE FAFESHMQEH VATMARQYVR 660 SIISSVQRVA LALSPSNISS QAGLRTPLGT PEALTLARWI CHSYRCYLGV DLLKSSSEGS 720 ESILKNLWHH TDAVMCCSLK ALPVFTFANQ AGLDMLETTL VALQDITLEK IFDDHGRKAL 780 FAEFPQIMQQ GFACLQGGIC LSSMGRPVSY ERAVAWKVLN EEETAHCICF MFINWSFV* |
| Expression -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| Uniprot | TISSUE SPECIFICITY: Highly expressed the developing vascular elements and the adaxial portion of cotyledons. Expressed in developing ovules, stamens and carpels. Expressed in procambium and shoot meristem. {ECO:0000269|PubMed:14701930, ECO:0000269|PubMed:15705957}. | |||||
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Probable transcription factor involved in the regulation of meristem development to promote lateral organ formation. May regulates procambial and vascular tissue formation or maintenance, and vascular development in inflorescence stems. {ECO:0000269|PubMed:15598805, ECO:0000269|PubMed:15705957, ECO:0000269|PubMed:16617092}. | |||||
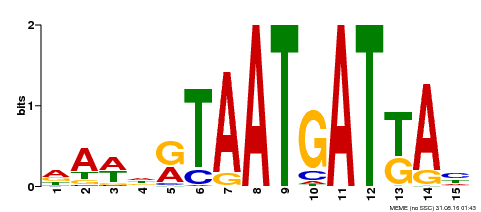
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00200 | DAP | Transfer from AT1G52150 | Download |
| |||
| Regulation -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | INDUCTION: By auxin. Repressed by miR165 and miR166. {ECO:0000269|PubMed:15773855, ECO:0000269|PubMed:16033795, ECO:0000269|PubMed:16617092, ECO:0000269|PubMed:17237362}. | |||||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_006426893.1 | 0.0 | homeobox-leucine zipper protein ATHB-15 | ||||
| Refseq | XP_006465680.1 | 0.0 | homeobox-leucine zipper protein ATHB-15 | ||||
| Refseq | XP_015386901.1 | 0.0 | homeobox-leucine zipper protein ATHB-15 | ||||
| Swissprot | Q9ZU11 | 0.0 | ATB15_ARATH; Homeobox-leucine zipper protein ATHB-15 | ||||
| TrEMBL | V4RYP0 | 0.0 | V4RYP0_9ROSI; Uncharacterized protein | ||||
| STRING | XP_006465679.1 | 0.0 | (Citrus sinensis) | ||||
| STRING | XP_006426892.1 | 0.0 | (Citrus clementina) | ||||
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT1G52150.1 | 0.0 | HD-ZIP family protein | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | Ciclev10024892m |
| Entrez Gene | 18036028 |
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



