 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Cc11_g07170 | ||||||||
| Common Name | GSCOC_T00018091001 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; asterids; lamiids; Gentianales; Rubiaceae; Ixoroideae; Coffeeae; Coffea
|
||||||||
| Family | C2H2 | ||||||||
| Protein Properties | Length: 396aa MW: 45958.2 Da PI: 8.4869 | ||||||||
| Description | C2H2 family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | zf-C2H2 | 20 | 1.9e-06 | 64 | 85 | 2 | 23 |
EETTTTEEESSHHHHHHHHHHT CS
zf-C2H2 2 kCpdCgksFsrksnLkrHirtH 23
+C++Cg sF+ + +L++H+++H
Cc11_g07170 64 TCEHCGVSFRKPAYLRQHMQSH 85
6*******************99 PP
| |||||||
| 2 | zf-C2H2 | 15.7 | 4.3e-05 | 114 | 138 | 1 | 23 |
EEET..TTTEEESSHHHHHHHHHHT CS
zf-C2H2 1 ykCp..dCgksFsrksnLkrHirtH 23
+kC dC+ s++rk++L+rH+ H
Cc11_g07170 114 FKCLvaDCHASYRRKDHLTRHLLQH 138
899999***************9887 PP
| |||||||
| 3 | zf-C2H2 | 19.4 | 2.8e-06 | 182 | 206 | 1 | 23 |
EEET..TTTEEESSHHHHHHHHHHT CS
zf-C2H2 1 ykCp..dCgksFsrksnLkrHirtH 23
y+C+ Cgk+F+ s L+rH +H
Cc11_g07170 182 YVCTevGCGKVFKFASKLRRHEESH 206
99*******************9988 PP
| |||||||
| 4 | zf-C2H2 | 14.9 | 7.6e-05 | 273 | 297 | 2 | 23 |
EET..TTTEEESSHHHHHHHHHH.T CS
zf-C2H2 2 kCp..dCgksFsrksnLkrHirt.H 23
kC+ C sFs+ksnL++H + H
Cc11_g07170 273 KCSfeGCLLSFSNKSNLNQHVKAvH 297
688889**************99877 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| SMART | SM00355 | 29 | 19 | 41 | IPR015880 | Zinc finger, C2H2-like |
| PROSITE profile | PS50157 | 8.517 | 19 | 46 | IPR007087 | Zinc finger, C2H2 |
| SuperFamily | SSF57667 | 1.98E-6 | 61 | 138 | No hit | No description |
| SMART | SM00355 | 0.007 | 63 | 85 | IPR015880 | Zinc finger, C2H2-like |
| PROSITE profile | PS50157 | 11.032 | 63 | 90 | IPR007087 | Zinc finger, C2H2 |
| Gene3D | G3DSA:3.30.160.60 | 2.6E-5 | 64 | 86 | IPR013087 | Zinc finger C2H2-type/integrase DNA-binding domain |
| PROSITE pattern | PS00028 | 0 | 65 | 85 | IPR007087 | Zinc finger, C2H2 |
| Gene3D | G3DSA:3.30.160.60 | 9.8E-9 | 108 | 140 | IPR013087 | Zinc finger C2H2-type/integrase DNA-binding domain |
| SMART | SM00355 | 0.029 | 114 | 138 | IPR015880 | Zinc finger, C2H2-like |
| PROSITE profile | PS50157 | 9.39 | 114 | 143 | IPR007087 | Zinc finger, C2H2 |
| PROSITE pattern | PS00028 | 0 | 116 | 138 | IPR007087 | Zinc finger, C2H2 |
| Gene3D | G3DSA:3.30.160.60 | 1.1E-4 | 141 | 168 | IPR013087 | Zinc finger C2H2-type/integrase DNA-binding domain |
| SMART | SM00355 | 0.6 | 143 | 168 | IPR015880 | Zinc finger, C2H2-like |
| PROSITE profile | PS50157 | 9.078 | 143 | 173 | IPR007087 | Zinc finger, C2H2 |
| PROSITE pattern | PS00028 | 0 | 145 | 168 | IPR007087 | Zinc finger, C2H2 |
| SuperFamily | SSF57667 | 2.02E-5 | 177 | 206 | No hit | No description |
| Gene3D | G3DSA:3.30.160.60 | 6.3E-8 | 179 | 206 | IPR013087 | Zinc finger C2H2-type/integrase DNA-binding domain |
| SMART | SM00355 | 0.0032 | 182 | 206 | IPR015880 | Zinc finger, C2H2-like |
| PROSITE profile | PS50157 | 11.219 | 182 | 211 | IPR007087 | Zinc finger, C2H2 |
| PROSITE pattern | PS00028 | 0 | 184 | 206 | IPR007087 | Zinc finger, C2H2 |
| SMART | SM00355 | 7 | 214 | 239 | IPR015880 | Zinc finger, C2H2-like |
| PROSITE pattern | PS00028 | 0 | 216 | 239 | IPR007087 | Zinc finger, C2H2 |
| SMART | SM00355 | 80 | 242 | 263 | IPR015880 | Zinc finger, C2H2-like |
| Gene3D | G3DSA:3.30.160.60 | 3.0E-5 | 253 | 294 | IPR013087 | Zinc finger C2H2-type/integrase DNA-binding domain |
| SMART | SM00355 | 0.0041 | 272 | 297 | IPR015880 | Zinc finger, C2H2-like |
| PROSITE profile | PS50157 | 11.468 | 272 | 302 | IPR007087 | Zinc finger, C2H2 |
| PROSITE pattern | PS00028 | 0 | 274 | 297 | IPR007087 | Zinc finger, C2H2 |
| SuperFamily | SSF57667 | 5.53E-7 | 282 | 325 | No hit | No description |
| SMART | SM00355 | 15 | 303 | 329 | IPR015880 | Zinc finger, C2H2-like |
| PROSITE pattern | PS00028 | 0 | 305 | 329 | IPR007087 | Zinc finger, C2H2 |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0005730 | Cellular Component | nucleolus | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| GO:0008097 | Molecular Function | 5S rRNA binding | ||||
| GO:0046872 | Molecular Function | metal ion binding | ||||
| GO:0080084 | Molecular Function | 5S rDNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 396 aa Download sequence Send to blast |
MVEEKENKRP VIFRDIRRYY CEYCGICRSK KSLISSHILS HHQEELKQKE EENGQAKDEP 60 KMNTCEHCGV SFRKPAYLRQ HMQSHSFEYQ VGGPPEKILQ ASLFHAVFSY ERPFKCLVAD 120 CHASYRRKDH LTRHLLQHQG KLFECPLDSC KCGFAYQGNM KRHVKEFHYD SSSNDVTSPK 180 QYVCTEVGCG KVFKFASKLR RHEESHVKLD TVEAFCAEPG CMKYFTNGEC LKEHVRSCHQ 240 YVICETCGTR QLKKNFKRHL CSHEAGCSQS KVKCSFEGCL LSFSNKSNLN QHVKAVHLGL 300 QPFSCRFPSC GMTFSFKHVR DNHEKSGRHV FTRGDFEESD EQFRSRPRGG RKRQYPVIET 360 LMRKRIVPPC ESDPILNEAS DYLSRFFSTE SEDEL* |
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Essential protein (PubMed:22353599). Isoform 1 is a transcription activator the binds both 5S rDNA and 5S rRNA and stimulates the transcription of 5S rRNA gene (PubMed:12711688, PubMed:22353599). Isoform 1 regulates 5S rRNA levels during development (PubMed:22353599). {ECO:0000269|PubMed:12711688, ECO:0000269|PubMed:22353599}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00229 | DAP | Transfer from AT1G72050 | Download |
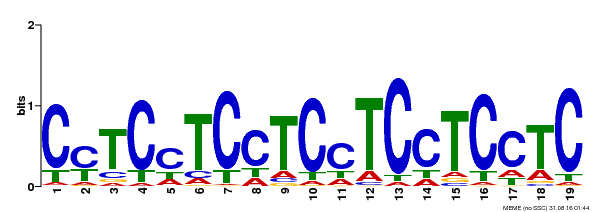
| |||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_027095861.1 | 0.0 | transcription factor IIIA-like | ||||
| Refseq | XP_027095862.1 | 0.0 | transcription factor IIIA-like | ||||
| Swissprot | Q84MZ4 | 1e-130 | TF3A_ARATH; Transcription factor IIIA | ||||
| TrEMBL | A0A068V6C1 | 0.0 | A0A068V6C1_COFCA; Uncharacterized protein | ||||
| STRING | XP_009622481.1 | 0.0 | (Nicotiana tomentosiformis) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Asterids | OGEA3644 | 24 | 47 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT1G72050.2 | 1e-120 | transcription factor IIIA | ||||



