 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Cc00_g08780 | ||||||||
| Common Name | GSCOC_T00007633001 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; asterids; lamiids; Gentianales; Rubiaceae; Ixoroideae; Coffeeae; Coffea
|
||||||||
| Family | C2H2 | ||||||||
| Protein Properties | Length: 323aa MW: 36030 Da PI: 5.2507 | ||||||||
| Description | C2H2 family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | zf-C2H2 | 21.8 | 4.8e-07 | 251 | 272 | 2 | 23 |
EETTTTEEESSHHHHHHHHHHT CS
zf-C2H2 2 kCpdCgksFsrksnLkrHirtH 23
C++Cgk F+r nL+ H+r H
Cc00_g08780 251 FCTICGKGFKRDANLRMHMRGH 272
6*******************98 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| SuperFamily | SSF57667 | 1.27E-5 | 248 | 275 | No hit | No description |
| PROSITE profile | PS50157 | 12.03 | 250 | 277 | IPR007087 | Zinc finger, C2H2 |
| SMART | SM00355 | 0.0026 | 250 | 272 | IPR015880 | Zinc finger, C2H2-like |
| Pfam | PF12874 | 5.8E-4 | 251 | 272 | No hit | No description |
| Gene3D | G3DSA:3.30.160.60 | 1.1E-5 | 251 | 276 | IPR013087 | Zinc finger C2H2-type/integrase DNA-binding domain |
| PROSITE pattern | PS00028 | 0 | 252 | 272 | IPR007087 | Zinc finger, C2H2 |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0006355 | Biological Process | regulation of transcription, DNA-templated | ||||
| GO:0010044 | Biological Process | response to aluminum ion | ||||
| GO:0010447 | Biological Process | response to acidic pH | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003676 | Molecular Function | nucleic acid binding | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| GO:0046872 | Molecular Function | metal ion binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 323 aa Download sequence Send to blast |
MDPDERLRTN PWTKSSSSAS DALRQLSPDN QSFTNFSSQQ QKWEDSSSMD YRTRIEQQFS 60 EFTNKRMRET SNSNNVQDWD PRGMLNNLSF LEQKIHQLQD LVHLIVGRRC QAVGQSNELL 120 VQQQQLITAD LTSIIVQLIS TAGSLLPSVK HSLSSVNPPV TQLGQFGGSI TPPVTSFDNS 180 GRVVHCDVKK VEDQPNEVDL IGNAGIEQNY VVEEHESKDE EDADEGENLP PGSYEILQLE 240 KEEILAPHTH FCTICGKGFK RDANLRMHMR GHGDEYKTPA ALAKPNKEST SEPTLIKRIT 300 IRELTATRAT LVAGATQRNS QS* |
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Probable transcription factor. Together with STOP2, plays a critical role in tolerance to major stress factors in acid soils such as proton H(+) and aluminum ion Al(3+). Required for the expression of genes in response to acidic stress (e.g. ALMT1 and MATE), and Al-activated citrate exudation. {ECO:0000269|PubMed:17535918, ECO:0000269|PubMed:18826429, ECO:0000269|PubMed:19321711, ECO:0000269|PubMed:23935008}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
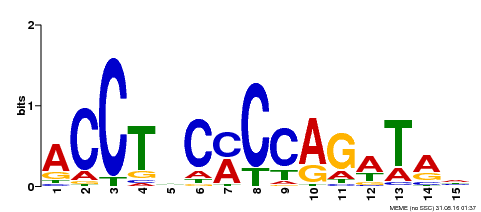
| Motif ID | Method | Source | Motif file |
| MP00196 | ampDAP | Transfer from AT1G34370 | Download |
| |||
| Regulation -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | INDUCTION: By shock H(+) and Al(3+) treatments. {ECO:0000269|PubMed:17535918}. | |||||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_027094878.1 | 0.0 | protein SENSITIVE TO PROTON RHIZOTOXICITY 1-like isoform X2 | ||||
| Refseq | XP_027094902.1 | 0.0 | protein SENSITIVE TO PROTON RHIZOTOXICITY 1-like isoform X2 | ||||
| Swissprot | Q9C8N5 | 1e-104 | STOP1_ARATH; Protein SENSITIVE TO PROTON RHIZOTOXICITY 1 | ||||
| TrEMBL | A0A068VI21 | 0.0 | A0A068VI21_COFCA; Uncharacterized protein | ||||
| STRING | XP_008237278.1 | 1e-138 | (Prunus mume) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Asterids | OGEA4754 | 24 | 41 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT1G34370.2 | 1e-101 | C2H2 family protein | ||||



