 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Carubv10028430m | ||||||||
| Common Name | CARUB_v10028430mg | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; malvids; Brassicales; Brassicaceae; Camelineae; Capsella
|
||||||||
| Family | bHLH | ||||||||
| Protein Properties | Length: 475aa MW: 53493.9 Da PI: 8.2019 | ||||||||
| Description | bHLH family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | HLH | 39.1 | 1.3e-12 | 306 | 351 | 4 | 54 |
HHHHHHHHHHHHHHHHHHHHHCTSCCC...TTS-STCHHHHHHHHHHHHHH CS
HLH 4 ahnerErrRRdriNsafeeLrellPkaskapskKlsKaeiLekAveYIksL 54
+h e+Er RR+++N++f Lr ++P+ K++K + Le Av+YI++L
Carubv10028430m 306 NHVEAERMRREKLNNRFYALRAVVPNV-----SKMDKTSLLEDAVRYINEL 351
799***********************6.....5***************998 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| Pfam | PF14215 | 3.4E-47 | 36 | 232 | IPR025610 | Transcription factor MYC/MYB N-terminal |
| SuperFamily | SSF47459 | 8.9E-17 | 300 | 366 | IPR011598 | Myc-type, basic helix-loop-helix (bHLH) domain |
| CDD | cd00083 | 1.96E-13 | 301 | 352 | No hit | No description |
| PROSITE profile | PS50888 | 16.818 | 302 | 351 | IPR011598 | Myc-type, basic helix-loop-helix (bHLH) domain |
| Pfam | PF00010 | 5.4E-10 | 306 | 351 | IPR011598 | Myc-type, basic helix-loop-helix (bHLH) domain |
| Gene3D | G3DSA:4.10.280.10 | 2.3E-15 | 306 | 372 | IPR011598 | Myc-type, basic helix-loop-helix (bHLH) domain |
| SMART | SM00353 | 6.1E-14 | 308 | 357 | IPR011598 | Myc-type, basic helix-loop-helix (bHLH) domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0042538 | Biological Process | hyperosmotic salinity response | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| GO:0005509 | Molecular Function | calcium ion binding | ||||
| GO:0046983 | Molecular Function | protein dimerization activity | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 475 aa Download sequence Send to blast |
MNNDENLSMI EALLTSDVWP PPPLPPANNL SLETTLQKRL HNVLNGTHVL WTYAVFWKPS 60 YNLLSGDSVL KWGDGVYNGG DEEKNGRRMR RRKKKTVPSS PEGKERRSKV LRELNSMISG 120 EPFPLVEVDD VNDDDDDDVE VTDTEWYYLV SMTWSSCNGS GLAGKAFDTC NPVWVTGLDQ 180 IYGSGCDRAK QGGGLGLQTI VCIPLDNGVL ELGSTELIRH NSDLFNKIRF LFHFEGSKDL 240 SGAPTSTQFR SRIHNPNTNP SPVYQPIQMN ISGEEFNRIN ATVIPEKKPG KKRGRKPNHG 300 REKPLNHVEA ERMRREKLNN RFYALRAVVP NVSKMDKTSL LEDAVRYINE LKSKAENSES 360 GKTAVEIQLK ELKKVMELQN ATSSSVCKDK EKKLSELKME VKVMGSDVMI RVESGKRNHP 420 GARFMNALMD LELEVNHTSI SVMNDLMIQQ ATVKMGLRIY EQEQLRDMLI SKIS* |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 4rqw_A | 9e-43 | 33 | 234 | 6 | 194 | Transcription factor MYC3 |
| 4rqw_B | 9e-43 | 33 | 234 | 6 | 194 | Transcription factor MYC3 |
| 4rs9_A | 9e-43 | 33 | 234 | 6 | 194 | Transcription factor MYC3 |
| 4yz6_A | 9e-43 | 33 | 234 | 6 | 194 | Transcription factor MYC3 |
| Search in ModeBase | ||||||
| Nucleic Localization Signal ? help Back to Top | |||
|---|---|---|---|
| No. | Start | End | Sequence |
| 1 | 86 | 91 | RRMRRR |
| 2 | 86 | 94 | RRMRRRKKK |
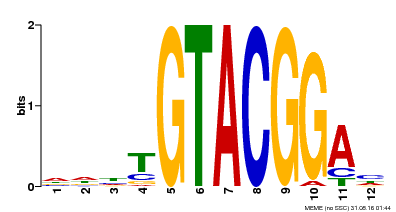
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00547 | DAP | Transfer from AT5G46830 | Download |
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | Carubv10028430m |
| Regulation -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | INDUCTION: By UV treatment. {ECO:0000269|PubMed:12679534}. | |||||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_006279851.1 | 0.0 | transcription factor bHLH28 | ||||
| Swissprot | Q9LUK7 | 0.0 | BH028_ARATH; Transcription factor bHLH28 | ||||
| TrEMBL | R0G888 | 0.0 | R0G888_9BRAS; Uncharacterized protein | ||||
| STRING | XP_006279851.1 | 0.0 | (Capsella rubella) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Malvids | OGEM13953 | 16 | 24 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT5G46830.1 | 0.0 | NACL-inducible gene 1 | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | Carubv10028430m |
| Entrez Gene | 17877237 |
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



