 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Carubv10026697m | ||||||||
| Common Name | CARUB_v10026697mg | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; malvids; Brassicales; Brassicaceae; Camelineae; Capsella
|
||||||||
| Family | NAC | ||||||||
| Protein Properties | Length: 350aa MW: 39791.8 Da PI: 5.0089 | ||||||||
| Description | NAC family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | NAM | 175.1 | 2e-54 | 8 | 136 | 2 | 129 |
NAM 2 ppGfrFhPtdeelvveyLkkkvegkkleleevikevdiykvePwdLpk...kvkaeekewyfFskrdkkyatgkrknratksgyWkatgkdkev 92
ppG+rFhPtdeelv++yLk+kv+ +++ +vik+vd+yk+ePwd+++ + ++ee+ewyfFs++dkky+tg+r+nrat sg+Wkatg+dk++
Carubv10026697m 8 PPGYRFHPTDEELVDYYLKNKVAYPGMQV-DVIKDVDLYKIEPWDIQElcgRGTGEEREWYFFSHKDKKYPTGTRTNRATGSGFWKATGRDKAI 100
9****************************.99**************954555555788************************************ PP
NAM 93 lskkgelvglkktLvfykgrapkgektdWvmheyrle 129
+s k+elvg++ktLvfykgrap+g+k+dW+mheyrle
Carubv10026697m 101 YS-KQELVGMRKTLVFYKGRAPNGQKSDWIMHEYRLE 136
**.999*****************************85 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| SuperFamily | SSF101941 | 3.01E-60 | 5 | 156 | IPR003441 | NAC domain |
| PROSITE profile | PS51005 | 58.378 | 7 | 156 | IPR003441 | NAC domain |
| Pfam | PF02365 | 3.4E-28 | 8 | 135 | IPR003441 | NAC domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0009620 | Biological Process | response to fungus | ||||
| GO:0009733 | Biological Process | response to auxin | ||||
| GO:0009735 | Biological Process | response to cytokinin | ||||
| GO:0009737 | Biological Process | response to abscisic acid | ||||
| GO:0009741 | Biological Process | response to brassinosteroid | ||||
| GO:0010981 | Biological Process | regulation of cell wall macromolecule metabolic process | ||||
| GO:0043068 | Biological Process | positive regulation of programmed cell death | ||||
| GO:0045491 | Biological Process | xylan metabolic process | ||||
| GO:0045893 | Biological Process | positive regulation of transcription, DNA-templated | ||||
| GO:0048759 | Biological Process | xylem vessel member cell differentiation | ||||
| GO:0090058 | Biological Process | metaxylem development | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0009531 | Cellular Component | secondary cell wall | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| GO:0042803 | Molecular Function | protein homodimerization activity | ||||
| GO:0043565 | Molecular Function | sequence-specific DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 350 aa Download sequence Send to blast |
MESLSHIPPG YRFHPTDEEL VDYYLKNKVA YPGMQVDVIK DVDLYKIEPW DIQELCGRGT 60 GEEREWYFFS HKDKKYPTGT RTNRATGSGF WKATGRDKAI YSKQELVGMR KTLVFYKGRA 120 PNGQKSDWIM HEYRLETDEN GPPHEEGWVV CRAFKKKLTT MNYNNPRTMM GSSAGQESNW 180 FTQQMDVANG NYYHLPDLES PRMFQGSSSS SLSSLHQNDQ DPYGVVLSTI NANPTTIMQR 240 DDGHVMSNDD DHMIMMNTST GGHHQSGLLV NDDHNDQVMD WQTLDKFVAS QLIMSQEEEE 300 VNKDPSDNSS NETFHHLSEE QATMVSINGN ASSSSSPCSF YSWAQNTQT* |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 3ulx_A | 4e-50 | 2 | 157 | 10 | 169 | Stress-induced transcription factor NAC1 |
| Search in ModeBase | ||||||
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Transcription activator that binds to the secondary wall NAC binding element (SNBE), 5'-(T/A)NN(C/T)(T/C/G)TNNNNNNNA(A/C)GN(A/C/T)(A/T)-3', and to the tracheary elements (TE) specific regulating cis-element (TERE), 5'-CTTNAAAGCNA-3', in the promoter of target genes (e.g. genes involved in secondary wall biosynthesis, cell wall modification such as xylan accumulation, and programmed cell death) (PubMed:20935069, PubMed:20952636, PubMed:20488898). Involved in xylem formation in roots and shoots, especially regulating metaxylem vessel differentiation by promoting immature xylem vessel-specific genes expression, especially genes regulating programmed cell death (PCD) and secondary wall formation in tracheary elements (TE) (PubMed:16103214, PubMed:20952636, PubMed:20488898). Can activate MYB25, MYB46, MYB58, MYB63, MYB83, MYB103, CESA4, LBD15, LBD30, ERF115, XCP1, XCP2, NAC010/SND3, KNAT7, ASL19 and ASL20 expression (PubMed:17890373, PubMed:18952777, PubMed:19088331, PubMed:19122102, PubMed:19808805, PubMed:20935069, PubMed:20952636). {ECO:0000269|PubMed:16103214, ECO:0000269|PubMed:17890373, ECO:0000269|PubMed:18952777, ECO:0000269|PubMed:19088331, ECO:0000269|PubMed:19122102, ECO:0000269|PubMed:19808805, ECO:0000269|PubMed:20488898, ECO:0000269|PubMed:20935069, ECO:0000269|PubMed:20952636}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00578 | DAP | Transfer from AT5G62380 | Download |
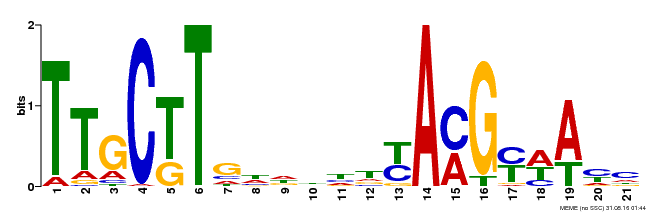
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | Carubv10026697m |
| Regulation -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | INDUCTION: By brassinosteroids (e.g. brassinolide BL), auxin (e.g. 2,4-dichlorphenoxyacetic acid 2,4-D) and cytokinin (e.g. kinetin), with a synergistic effect (PubMed:16103214). Accumulates during infection by the soilborne fungal pathogen Verticillium longisporum, especially in tissues undergoing de novo xylem formation (PubMed:23023171). {ECO:0000269|PubMed:16103214, ECO:0000269|PubMed:23023171}. | |||||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | BT026510 | 0.0 | BT026510.1 Arabidopsis thaliana At5g62380 mRNA, complete cds. | |||
| GenBank | DQ056734 | 0.0 | DQ056734.1 Arabidopsis thaliana no apical meristem family protein (At5g62380) mRNA, complete cds. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_006280728.1 | 0.0 | NAC domain-containing protein 101 | ||||
| Swissprot | Q9LVA1 | 0.0 | NC101_ARATH; NAC domain-containing protein 101 | ||||
| TrEMBL | R0EXL1 | 0.0 | R0EXL1_9BRAS; Uncharacterized protein | ||||
| STRING | Cagra.4003s0027.1.p | 0.0 | (Capsella grandiflora) | ||||
| STRING | XP_006280728.1 | 0.0 | (Capsella rubella) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Malvids | OGEM12248 | 17 | 32 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT5G62380.1 | 0.0 | NAC-domain protein 101 | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | Carubv10026697m |
| Entrez Gene | 17877287 |
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



