 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Cagra.0094s0026.1.p | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; malvids; Brassicales; Brassicaceae; Camelineae; Capsella
|
||||||||
| Family | HD-ZIP | ||||||||
| Protein Properties | Length: 685aa MW: 76165.4 Da PI: 6.1796 | ||||||||
| Description | HD-ZIP family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | Homeobox | 57.8 | 1.8e-18 | 62 | 117 | 1 | 56 |
TT--SS--HHHHHHHHHHHHHSSS--HHHHHHHHHHCTS-HHHHHHHHHHHHHHHH CS
Homeobox 1 rrkRttftkeqleeLeelFeknrypsaeereeLAkklgLterqVkvWFqNrRakek 56
r k +++t++q++eLe++F+ +++p++++r eL kkl L+ +q+k+WFqNrR+++k
Cagra.0094s0026.1.p 62 RTKYHRHTSYQIQELESFFKVCPHPNEKQRLELGKKLTLESKQIKFWFQNRRTQMK 117
3455678999*******************************************999 PP
| |||||||
| 2 | START | 146.6 | 2.4e-46 | 228 | 432 | 2 | 206 |
HHHHHHHHHHHHHHHC-TT-EEEEEXCCTTEEEEEEESSS.SCEEEEEEEECCSCHHHHHHHHHCCCGGCT-TT-S....EEEEEEEECT CS
START 2 laeeaaqelvkkalaeepgWvkssesengdevlqkfeeskvdsgealrasgvvdmvlallveellddkeqWdetla....kaetleviss 87
la ea++e++k+ +++ p+W + s ++e++ e +r+sg+v +++ lv l+d++ +W e+++ a+t++vis+
Cagra.0094s0026.1.p 228 LAMEAMEEFLKLEELDNPLWNSKS----EKESMNHNE-----YRSSSRESGLVLINSVALVDALMDTN-KWAEMFEcivaVASTVKVISN 307
5789****************9999....444444442.....22347*********************.********************* PP
T......EEEEEEEEXXTTXX-SSX.EEEEEEEEEEE.TTS-EEEEEEEEE-TTS--.-TTSEE-EESSEEEEEEEECTCEEEEEEEE-E CS
START 88 g......galqlmvaelqalsplvp.RdfvfvRyirqlgagdwvivdvSvdseqkppesssvvRaellpSgiliepksnghskvtwvehv 170
g g lqlm+ae+q++splvp + f+Ry++q+g+g w++vdvS d +++ ++ +s+ ++++pSg++i++ +ng skvtw+eh
Cagra.0094s0026.1.p 308 GsdgsrnGSLQLMQAEFQVMSPLVPiKQKKFLRYCKQHGDGLWAVVDVSYDINREDENLKSYGGSKKFPSGCIIQDIGNGCSKVTWIEHL 397
*******************************************************999******************************** PP
E--SSXXHHHHHHHHHHHHHHHHHHHHHHTXXXXXX CS
START 171 dlkgrlphwllrslvksglaegaktwvatlqrqcek 206
+++++++ +++ +l++s++a ga +w+atlqrqce+
Cagra.0094s0026.1.p 398 EYEESHINSVY-QLLGSSVALGATKWLATLQRQCES 432
*******9998.689999****************95 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| Gene3D | G3DSA:1.10.10.60 | 3.6E-19 | 46 | 119 | IPR009057 | Homeodomain-like |
| SuperFamily | SSF46689 | 6.68E-18 | 55 | 119 | IPR009057 | Homeodomain-like |
| SMART | SM00389 | 5.7E-15 | 58 | 123 | IPR001356 | Homeobox domain |
| PROSITE profile | PS50071 | 16.86 | 59 | 119 | IPR001356 | Homeobox domain |
| CDD | cd00086 | 1.67E-15 | 62 | 119 | No hit | No description |
| Pfam | PF00046 | 3.2E-16 | 62 | 117 | IPR001356 | Homeobox domain |
| PROSITE profile | PS50848 | 39.207 | 218 | 435 | IPR002913 | START domain |
| CDD | cd08875 | 1.20E-100 | 223 | 431 | No hit | No description |
| SuperFamily | SSF55961 | 3.02E-27 | 224 | 433 | No hit | No description |
| SMART | SM00234 | 6.8E-33 | 227 | 432 | IPR002913 | START domain |
| Pfam | PF01852 | 7.0E-40 | 228 | 432 | IPR002913 | START domain |
| SuperFamily | SSF55961 | 1.1E-17 | 450 | 671 | No hit | No description |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0008289 | Molecular Function | lipid binding | ||||
| GO:0043565 | Molecular Function | sequence-specific DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 685 aa Download sequence Send to blast |
MNGDLDVDMS RGDLNPSFFH GRLKDDEFES RSLSDDDSFD AMSGDENKQE EQRPKKKKKK 60 KRTKYHRHTS YQIQELESFF KVCPHPNEKQ RLELGKKLTL ESKQIKFWFQ NRRTQMKTQL 120 ERHENVILRQ ENEKLRVENG FLKESMRGSL CIDCGGAVIP GEVSFEQHQL RIENAKLKDE 180 LDRICALANR FIGGSISLEQ PSNGGIGSQH FPIGNGFSGG TSQMFMDLAM EAMEEFLKLE 240 ELDNPLWNSK SEKESMNHNE YRSSSRESGL VLINSVALVD ALMDTNKWAE MFECIVAVAS 300 TVKVISNGSD GSRNGSLQLM QAEFQVMSPL VPIKQKKFLR YCKQHGDGLW AVVDVSYDIN 360 REDENLKSYG GSKKFPSGCI IQDIGNGCSK VTWIEHLEYE ESHINSVYQL LGSSVALGAT 420 KWLATLQRQC ESFTSLLSSQ DHTGLSLAGT KSILKLAQRM KLNFYSGITA SSVHKWEKLN 480 AENVGQDTRI LTRKSFEPSG IVLSAATSLW LPVTQQRLFE FLCDGKCRNQ WDILSNGASM 540 EITLLVPKGQ QEGSCVSLLC AAGKDQNESS MLILQETWND ASGALVVYAP VDFPSMNVVM 600 SGGDSGYVAL LPSGFSILPD GSSLSDQIDT NGNQESKGCL LTVGFQILVN SLPTAKLNVE 660 SVETVNNLIA CTIHKIRAAL RIPA* |
| Nucleic Localization Signal ? help Back to Top | |||
|---|---|---|---|
| No. | Start | End | Sequence |
| 1 | 55 | 61 | KKKKKKR |
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
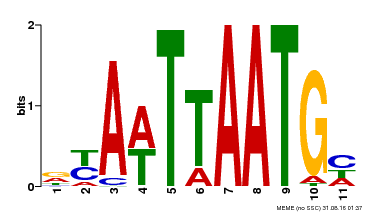
| UniProt | Probable transcription factor that binds to the DNA sequence 5'-GCATTAAATGC-3'. {ECO:0000269|PubMed:16778018}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00559 | DAP | Transfer from AT5G52170 | Download |
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | Cagra.0094s0026.1.p |
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | AB025603 | 1e-157 | AB025603.1 Arabidopsis thaliana genomic DNA, chromosome 5, BAC clone:F17P19. | |||
| GenBank | CP002688 | 1e-157 | CP002688.1 Arabidopsis thaliana chromosome 5 sequence. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_006282099.1 | 0.0 | homeobox-leucine zipper protein HDG7 | ||||
| Swissprot | Q9LTK3 | 0.0 | HDG7_ARATH; Homeobox-leucine zipper protein HDG7 | ||||
| TrEMBL | R0F0C9 | 0.0 | R0F0C9_9BRAS; Uncharacterized protein | ||||
| STRING | Cagra.0094s0026.1.p | 0.0 | (Capsella grandiflora) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Malvids | OGEM1128 | 27 | 105 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT5G52170.1 | 0.0 | homeodomain GLABROUS 7 | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | Cagra.0094s0026.1.p |
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



