 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | CCG017460.1 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; fabids; Malpighiales; Salicaceae; Saliceae; Populus
|
||||||||
| Family | bZIP | ||||||||
| Protein Properties | Length: 423aa MW: 46485.4 Da PI: 10.7778 | ||||||||
| Description | bZIP family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | bZIP_1 | 44 | 4.8e-14 | 345 | 391 | 5 | 51 |
CHHHCHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHH CS
bZIP_1 5 krerrkqkNReAArrsRqRKkaeieeLeekvkeLeaeNkaLkkelee 51
+r+rr++kNRe+A rsR+RK+a++ eLe++v++L++ Nk+L+ + e
CCG017460.1 345 RRQRRMIKNRESAARSRARKQAYTLELEDEVAKLKERNKELQRKQAE 391
79************************************999866544 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| SMART | SM00338 | 1.3E-12 | 341 | 406 | IPR004827 | Basic-leucine zipper domain |
| PROSITE profile | PS50217 | 10.932 | 343 | 392 | IPR004827 | Basic-leucine zipper domain |
| CDD | cd14707 | 1.71E-23 | 345 | 399 | No hit | No description |
| Pfam | PF00170 | 2.0E-12 | 345 | 392 | IPR004827 | Basic-leucine zipper domain |
| Gene3D | G3DSA:1.20.5.170 | 7.0E-14 | 345 | 392 | No hit | No description |
| SuperFamily | SSF57959 | 9.14E-11 | 345 | 393 | No hit | No description |
| PROSITE pattern | PS00036 | 0 | 348 | 363 | IPR004827 | Basic-leucine zipper domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0009414 | Biological Process | response to water deprivation | ||||
| GO:0009651 | Biological Process | response to salt stress | ||||
| GO:0009738 | Biological Process | abscisic acid-activated signaling pathway | ||||
| GO:0045893 | Biological Process | positive regulation of transcription, DNA-templated | ||||
| GO:0000976 | Molecular Function | transcription regulatory region sequence-specific DNA binding | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 423 aa Download sequence Send to blast |
MDSHWDFKNF GNASPGLGSV RKSPENPPLV RQSSVYSLTF DEFQNTWGGG LEKDFGSMNM 60 DELLKNIWTA EETQAMTNTV GVGGEGSTPD GNLQRQGSLT LPRTLSQKTV DEVWRDLIKE 120 TSGVAGSNLP QRQQTLREMT LEEFLVRAGV VREDTQQIGR PNSSGLFSEL SQLNNNNNTT 180 GLALGFQQPN GNNGLMGTGI MKNNLVSNQP SRLALNVGGI RPSQQLPQPQ QQQQQPLFPK 240 PAATVVFASP LHVANNARLA SPGVRGPVVG IADRSVNNGL AHSRGMEMVS LAARGVTVAT 300 GSPANRISPD VIAKSNADTS SLSPVPFVFS RGRKPSAALE KVVERRQRRM IKNRESAARS 360 RARKQAYTLE LEDEVAKLKE RNKELQRKQA EIFEMQKNQF LETMKAQWGG KRQCLRRTLT 420 GPW |
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Binds to the ABA-responsive element (ABRE). Mediates stress-responsive ABA signaling. {ECO:0000269|PubMed:11884679, ECO:0000269|PubMed:15361142}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00038 | PBM | Transfer from AT4G34000 | Download |
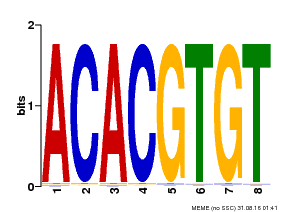
| |||
| Regulation -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | INDUCTION: Up-regulated by drought, salt, abscisic acid (ABA). {ECO:0000269|PubMed:10636868, ECO:0000269|PubMed:16284313}. | |||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | EF405964 | 0.0 | EF405964.1 Populus trichocarpa abscisic acid responsive elements-binding protein 2 (ABF2-1) mRNA, complete cds. | |||
| GenBank | EF405965 | 0.0 | EF405965.1 Populus trichocarpa abscisic acid responsive elements-binding protein 2 (ABF2-2) mRNA, complete cds. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_011000551.1 | 0.0 | PREDICTED: ABSCISIC ACID-INSENSITIVE 5-like protein 7 | ||||
| Refseq | XP_011000560.1 | 0.0 | PREDICTED: ABSCISIC ACID-INSENSITIVE 5-like protein 7 | ||||
| Refseq | XP_011000568.1 | 0.0 | PREDICTED: ABSCISIC ACID-INSENSITIVE 5-like protein 7 | ||||
| Refseq | XP_011000576.1 | 0.0 | PREDICTED: ABSCISIC ACID-INSENSITIVE 5-like protein 7 | ||||
| Swissprot | Q9M7Q3 | 1e-116 | AI5L6_ARATH; ABSCISIC ACID-INSENSITIVE 5-like protein 6 | ||||
| TrEMBL | A3FM74 | 0.0 | A3FM74_POPTR; Abscisic acid responsive elements-binding protein 2 | ||||
| STRING | POPTR_0009s10400.1 | 0.0 | (Populus trichocarpa) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Fabids | OGEF1962 | 34 | 81 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT4G34000.2 | 1e-117 | abscisic acid responsive elements-binding factor 3 | ||||



