 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | CCG013253.3 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; fabids; Malpighiales; Salicaceae; Saliceae; Populus
|
||||||||
| Family | bHLH | ||||||||
| Protein Properties | Length: 565aa MW: 62222.8 Da PI: 6.5106 | ||||||||
| Description | bHLH family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | HLH | 39.8 | 8e-13 | 342 | 420 | 4 | 55 |
HHHHHHHHHHHHHHHHHHHHHCTSCCC..............................TTS-STCHHHHHHHHHHHHHHH CS
HLH 4 ahnerErrRRdriNsafeeLrellPkas...........................kapskKlsKaeiLekAveYIksLq 55
hn ErrRRdriN+++ L+el+P+++ +Ka++L +A+eY+ksLq
CCG013253.3 342 VHNLSERRRRDRINEKMRALQELIPHCYkeysielillqafilllpkcpslyevkLVLMLQTDKASMLDEAIEYLKSLQ 420
6*************************999********************99999733444479***************9 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| PROSITE profile | PS50888 | 16.533 | 338 | 419 | IPR011598 | Myc-type, basic helix-loop-helix (bHLH) domain |
| CDD | cd00083 | 2.80E-13 | 341 | 424 | No hit | No description |
| Gene3D | G3DSA:4.10.280.10 | 3.1E-17 | 342 | 368 | IPR011598 | Myc-type, basic helix-loop-helix (bHLH) domain |
| SuperFamily | SSF47459 | 1.18E-15 | 342 | 370 | IPR011598 | Myc-type, basic helix-loop-helix (bHLH) domain |
| Pfam | PF00010 | 7.7E-10 | 342 | 420 | IPR011598 | Myc-type, basic helix-loop-helix (bHLH) domain |
| SMART | SM00353 | 5.9E-12 | 344 | 425 | IPR011598 | Myc-type, basic helix-loop-helix (bHLH) domain |
| Gene3D | G3DSA:4.10.280.10 | 3.1E-17 | 400 | 428 | IPR011598 | Myc-type, basic helix-loop-helix (bHLH) domain |
| SuperFamily | SSF47459 | 1.18E-15 | 401 | 435 | IPR011598 | Myc-type, basic helix-loop-helix (bHLH) domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0009704 | Biological Process | de-etiolation | ||||
| GO:0010161 | Biological Process | red light signaling pathway | ||||
| GO:0010244 | Biological Process | response to low fluence blue light stimulus by blue low-fluence system | ||||
| GO:0010600 | Biological Process | regulation of auxin biosynthetic process | ||||
| GO:0010928 | Biological Process | regulation of auxin mediated signaling pathway | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| GO:0046983 | Molecular Function | protein dimerization activity | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 565 aa Download sequence Send to blast |
MNHYIPDWNF EGDLPVSNQK KPIELGNDLV ELLWRNGQVV LHSQAHRKPS PHVQKHDSPT 60 VKGSGSILNS SHLIQDDDAV SWIQYPLEDS FEKEFCSNFF SELPPPLSDQ ILEEKSAKFD 120 ASAPSQQQHQ HQHQRQLNNN KPHVVSEFSG NPMPPPRIQV PEKNHAGVGG FGEAVNANFS 180 QFSAPFKGGD FRTSNGQFGG QGSGNSPQGE VRECSVVTVG SSNQIPHDRD MSRASSNAVG 240 TSTAFSTGPS MDDPRKIVSE SERGKTETLE ATLTSSSGGS GSSFGRTCKQ SAGPSGSQKR 300 KTIDTEDSEY QSEAAELDLD SMAGNNPTKR SGSTRRNRAA EVHNLSERRR RDRINEKMRA 360 LQELIPHCYK EYSIELILLQ AFILLLPKCP SLYEVKLVLM LQTDKASMLD EAIEYLKSLQ 420 LQLQVMWMCG GMAPMLFPGV QHFMSRMGMG PPLSSMQNPM HLPRVQLIDQ SISMAPTQNQ 480 GVMCQTPELN PVNFHNQMQN PAFADQYARF MGFHMQAASQ PMNMFRFGSQ TVQQNQMMAP 540 TSSVGGPLSA GTAVGEAPPS DDKMG |
| Nucleic Localization Signal ? help Back to Top | |||
|---|---|---|---|
| No. | Start | End | Sequence |
| 1 | 346 | 351 | ERRRRD |
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00606 | ChIP-seq | Transfer from AT2G43010 | Download |
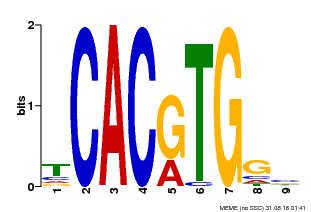
| |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_011032742.1 | 0.0 | PREDICTED: transcription factor PIF4-like isoform X2 | ||||
| TrEMBL | B9GSQ9 | 0.0 | B9GSQ9_POPTR; Uncharacterized protein | ||||
| STRING | POPTR_0005s22870.1 | 0.0 | (Populus trichocarpa) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Fabids | OGEF8146 | 29 | 44 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT2G43010.2 | 1e-43 | phytochrome interacting factor 4 | ||||



