 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | CCG002418.1 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; fabids; Malpighiales; Salicaceae; Saliceae; Populus
|
||||||||
| Family | MYB_related | ||||||||
| Protein Properties | Length: 486aa MW: 53727.7 Da PI: 5.9509 | ||||||||
| Description | MYB_related family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | Myb_DNA-binding | 52.1 | 1.5e-16 | 59 | 103 | 1 | 47 |
TSSS-HHHHHHHHHHHHHTTTT-HHHHHHHHTTTS-HHHHHHHHHHH CS
Myb_DNA-binding 1 rgrWTteEdellvdavkqlGggtWktIartmgkgRtlkqcksrwqky 47
r rWT+eE++++++a k++G W++I +++g ++t+ q++s+ qk+
CCG002418.1 59 RERWTEEEHKKFLEALKLYGRA-WRRIEEHVG-TKTAVQIRSHAQKF 103
78******************88.*********.************98 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| SuperFamily | SSF46689 | 1.2E-16 | 53 | 107 | IPR009057 | Homeodomain-like |
| PROSITE profile | PS51294 | 21.476 | 54 | 108 | IPR017930 | Myb domain |
| TIGRFAMs | TIGR01557 | 5.7E-17 | 57 | 106 | IPR006447 | Myb domain, plants |
| SMART | SM00717 | 2.5E-13 | 58 | 106 | IPR001005 | SANT/Myb domain |
| Pfam | PF00249 | 5.7E-14 | 59 | 102 | IPR001005 | SANT/Myb domain |
| Gene3D | G3DSA:1.10.10.60 | 1.4E-9 | 59 | 99 | IPR009057 | Homeodomain-like |
| CDD | cd00167 | 3.24E-10 | 61 | 104 | No hit | No description |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0007623 | Biological Process | circadian rhythm | ||||
| GO:0009734 | Biological Process | auxin-activated signaling pathway | ||||
| GO:0010600 | Biological Process | regulation of auxin biosynthetic process | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 486 aa Download sequence Send to blast |
MQDECGGTRL NLVLPAGNGI SLSATLNNAS GQQIKEQFSC GSDFSSPKAR KPYTITKQRE 60 RWTEEEHKKF LEALKLYGRA WRRIEEHVGT KTAVQIRSHA QKFFSKVVRE SGDSNTSSVE 120 SIEIPPPRPK RKPMHPYPRK LAHPLEKELL IPEKSLRSSS PNFSISEQEN QSPTSVLSAV 180 GSDALGSTDS DTANHSLSPV SFAGGVHHAD SSPEEDGYPS PATASSVPDE QFPKVEKLDS 240 SPKENVSSEE PVVEETSTRS LKLFGWTVLV TECHKPSSPN VGTSKLSTLD TAEEKLVRPL 300 TLNNVAAEFP SRNGESTWSP LPHGSHGALC YMKFQKENSS PAQNDSATLP WWTFYGAMPF 360 PCIPFHKKEH AIENLDSKGD EVQDKEIPKE VSWTGSTSGS VSEGENGDKN MDAETESQQF 420 SYEEKEISPI FELKLTKKSA SSGSEVINEK CPKGFVPYKK RIAERDSQSS TITSEEREEQ 480 RIRLCL |
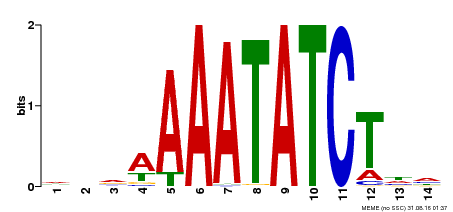
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00515 | DAP | Transfer from AT5G17300 | Download |
| |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_011035339.1 | 0.0 | PREDICTED: protein REVEILLE 1-like isoform X2 | ||||
| TrEMBL | A0A2K2AR85 | 0.0 | A0A2K2AR85_POPTR; Uncharacterized protein | ||||
| STRING | POPTR_0004s07270.1 | 0.0 | (Populus trichocarpa) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Fabids | OGEF2956 | 33 | 72 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT5G17300.1 | 6e-53 | MYB_related family protein | ||||



