 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | CA07g08600 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; asterids; lamiids; Solanales; Solanaceae; Solanoideae; Capsiceae; Capsicum
|
||||||||
| Family | FAR1 | ||||||||
| Protein Properties | Length: 866aa MW: 99414 Da PI: 7.2331 | ||||||||
| Description | FAR1 family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | FAR1 | 85 | 1.2e-26 | 90 | 192 | 2 | 91 |
FAR1 2 fYneYAkevGFsvrkskskkskrngeitkrtfvCskegkreeekkk.............tekerrtraetrtgCkaklkvkkekdgkwevtkleleHnH 87
fY+eYA+++GF++ +++s++sk+++e+++++f Cs++g++++ +k+ e++ +ra +t+Cka+++vk++ dgkw ++++e+eHnH
CA07g08600 90 FYQEYARSMGFNTAIQNSRRSKTSREFIDAKFACSRYGTKRDYEKSanrprsrqgnkqdPENATGRRACAKTDCKASMHVKRRPDGKWIIHRFEKEHNH 188
9**************************************9999999899999888776555556699******************************** PP
FAR1 88 elap 91
el p
CA07g08600 189 ELLP 192
*975 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| Pfam | PF03101 | 1.3E-24 | 90 | 192 | IPR004330 | FAR1 DNA binding domain |
| Pfam | PF10551 | 1.3E-25 | 290 | 382 | IPR018289 | MULE transposase domain |
| PROSITE profile | PS50966 | 9.458 | 570 | 606 | IPR007527 | Zinc finger, SWIM-type |
| Pfam | PF04434 | 0.0011 | 578 | 603 | IPR007527 | Zinc finger, SWIM-type |
| SMART | SM00575 | 6.6E-5 | 581 | 608 | IPR006564 | Zinc finger, PMZ-type |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0009585 | Biological Process | red, far-red light phototransduction | ||||
| GO:0010218 | Biological Process | response to far red light | ||||
| GO:0042753 | Biological Process | positive regulation of circadian rhythm | ||||
| GO:0045893 | Biological Process | positive regulation of transcription, DNA-templated | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| GO:0008270 | Molecular Function | zinc ion binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 866 aa Download sequence Send to blast |
MDIDLRLPSQ DHDKEEEEEQ NGIINMLDNE EKMHSDDGMH GMLVIEEKMH AEDGGDMNTP 60 IGTMIEFKED VNLEPLAGME FESHGEAYAF YQEYARSMGF NTAIQNSRRS KTSREFIDAK 120 FACSRYGTKR DYEKSANRPR SRQGNKQDPE NATGRRACAK TDCKASMHVK RRPDGKWIIH 180 RFEKEHNHEL LPAQAVSEQT RRMYAAMARQ FAEYKNVVGL KSDTKGPFEK GRNSAMEGGD 240 INVLLEFFIQ MQNLNSNFFY AVDVGEDQRV RNLFWVDAKA RHDYANFNDV VSFDTAYVRN 300 KYKMPLALFV GVNQHFQFML LGCALVSDES AATFFWVMRT WLKAMGGQAP KTVITDHDQV 360 LKSVIPEALP LSLHYFCLWH ILGKVSDTLN HIIKQNEKFM PKFEKCIHRS WTDEEFEKRW 420 RKLVDKFDLR EAELIHSLYE DRMKWAPTFI RDVLLAGMSM VLRSESVNSF FDKYVHKKTT 480 VQELVKQYES ILQDRYEEEA KADSDTWNKQ PALKSPSPFE KQVAGLYTNA VFKKFQAEVL 540 GAVACIPKRE KQDATTITYR VQDFEKTQEF IVTLDEMKSE ISCLCHLFEF KGYLCRHTLI 600 VLQICRVSSI PSQYILKRWT KDAKSKYSMT DGSEELQSRF QRYNELCHRA MKLSEEGSLS 660 QESYSFALRA LDDAFGSCVT FNNSNKNMLE AGTSSASGIL CIEDDNQSRS MSKTNKKKNN 720 FTKKRKVNSE PDVMAVGPAD SLQQMDKLNS RPVTLDGYFG PQQSVQGMVQ LNLMAPTRDN 780 YYGNQQTIQG LGQLNSIAPT HDGYYGAQPT MHGLVCGLLV LELLTQINPK SFLQFHFYCC 840 RGKWTFSALQ VSHMVFGLVS NILRL* |
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
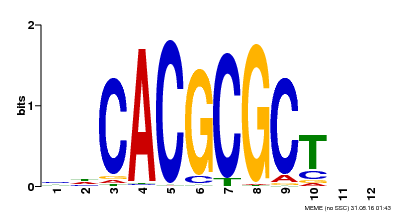
| UniProt | Transcription activator that recognizes and binds to the DNA consensus sequence 5'-CACGCGC-3'. Activates the expression of FHY1 and FHL involved in light responses. When associated with PHYA, protects it from being recognized and degraded by the COP1/SPA complex. Positive regulator of chlorophyll biosynthesis via the activation of HEMB1 gene expression. {ECO:0000269|PubMed:11889039, ECO:0000269|PubMed:12753585, ECO:0000269|PubMed:17012604, ECO:0000269|PubMed:18033885, ECO:0000269|PubMed:18715961, ECO:0000269|PubMed:22634759}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00078 | ChIP-seq | Transfer from AT3G22170 | Download |
| |||
| Regulation -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | INDUCTION: Down-regulated after exposure to far-red light. Subject to a negative feedback regulation by PHYA signaling. Up-regulated by white light. {ECO:0000269|PubMed:11889039, ECO:0000269|PubMed:18033885, ECO:0000269|PubMed:22634759}. | |||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | BT013715 | 0.0 | BT013715.1 Lycopersicon esculentum clone 132560F, mRNA sequence. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_016580413.1 | 0.0 | PREDICTED: LOW QUALITY PROTEIN: protein FAR-RED ELONGATED HYPOCOTYL 3-like | ||||
| Swissprot | Q9LIE5 | 0.0 | FHY3_ARATH; Protein FAR-RED ELONGATED HYPOCOTYL 3 | ||||
| TrEMBL | A0A2G2Z0V7 | 0.0 | A0A2G2Z0V7_CAPAN; Protein FAR-RED ELONGATED HYPOCOTYL 3 | ||||
| STRING | XP_009804946.1 | 0.0 | (Nicotiana sylvestris) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Asterids | OGEA496 | 21 | 103 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT3G22170.2 | 0.0 | far-red elongated hypocotyls 3 | ||||



