 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | C.cajan_12636 | ||||||||
| Common Name | KK1_013025 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; fabids; Fabales; Fabaceae; Papilionoideae; Phaseoleae; Cajanus
|
||||||||
| Family | BES1 | ||||||||
| Protein Properties | Length: 292aa MW: 31834.7 Da PI: 9.5089 | ||||||||
| Description | BES1 family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | DUF822 | 212.4 | 1.1e-65 | 5 | 145 | 1 | 148 |
DUF822 1 ggsgrkptwkErEnnkrRERrRRaiaakiyaGLRaqGnyklpkraDnneVlkALcreAGwvvedDGttyrkgskpleeaeaagssasaspesslqs 96
g++gr+ptwkErEnnkrRERrRRaiaakiyaGLRaqGnyklpk++DnneVlkALc+eAGw+ve+DGttyrkg+k++ + + +g + s+++s q
C.cajan_12636 5 GSTGRMPTWKERENNKRRERRRRAIAAKIYAGLRAQGNYKLPKHCDNNEVLKALCNEAGWMVEEDGTTYRKGCKRPASGSEIGGAMDSSIQASPQ- 99
589*************************************************************************7777777777766666655. PP
DUF822 97 slkssalaspvesysaspksssfpspssldsislasaasllpvlsvlslvss 148
ss+++spv+sy+asp+sssfpsp+++d +++ + l+p++++ +++++
C.cajan_12636 100 ---SSSFPSPVPSYQASPTSSSFPSPTRIDANNS---SFLIPFIRNITSIPA 145
...9**************************9977...488888888777665 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| Pfam | PF05687 | 3.6E-61 | 7 | 128 | IPR008540 | BES1/BZR1 plant transcription factor, N-terminal |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0009742 | Biological Process | brassinosteroid mediated signaling pathway | ||||
| GO:0045892 | Biological Process | negative regulation of transcription, DNA-templated | ||||
| GO:0048316 | Biological Process | seed development | ||||
| GO:0048481 | Biological Process | plant ovule development | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0005829 | Cellular Component | cytosol | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 292 aa Download sequence Send to blast |
MTGGGSTGRM PTWKERENNK RRERRRRAIA AKIYAGLRAQ GNYKLPKHCD NNEVLKALCN 60 EAGWMVEEDG TTYRKGCKRP ASGSEIGGAM DSSIQASPQS SSFPSPVPSY QASPTSSSFP 120 SPTRIDANNS SFLIPFIRNI TSIPANLPPL RISNSAPVTP PLSSPRASKR KANFHPLFAT 180 SAPSSPTRRH HLATSTIPEC DESDASTVDS ASGRWVSFQV QTMAAAPPSP TFNLMKPAMQ 240 IGSQDVNDGM QMQWERGSDF DFENGRVKPW EGERIHEVGM DDLELTLGVG KP |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 5zd4_A | 5e-26 | 9 | 88 | 372 | 449 | Maltose-binding periplasmic protein,Protein BRASSINAZOLE-RESISTANT 1 |
| 5zd4_B | 5e-26 | 9 | 88 | 372 | 449 | Maltose-binding periplasmic protein,Protein BRASSINAZOLE-RESISTANT 1 |
| 5zd4_C | 5e-26 | 9 | 88 | 372 | 449 | Maltose-binding periplasmic protein,Protein BRASSINAZOLE-RESISTANT 1 |
| 5zd4_D | 5e-26 | 9 | 88 | 372 | 449 | Maltose-binding periplasmic protein,Protein BRASSINAZOLE-RESISTANT 1 |
| Search in ModeBase | ||||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00237 | DAP | Transfer from AT1G75080 | Download |
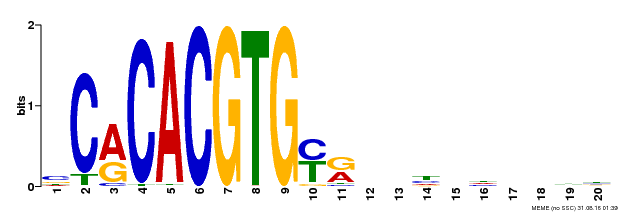
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | C.cajan_12636 |
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | EF533701 | 7e-90 | EF533701.1 Glycine max clone BAC GM_WBb095P01, complete sequence. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_029127596.1 | 0.0 | BES1/BZR1 homolog protein 2 | ||||
| Swissprot | Q94A43 | 3e-96 | BEH2_ARATH; BES1/BZR1 homolog protein 2 | ||||
| TrEMBL | A0A151TI63 | 0.0 | A0A151TI63_CAJCA; BES1/BZR1 isogeny protein 2 | ||||
| STRING | GLYMA01G38450.1 | 1e-137 | (Glycine max) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Fabids | OGEF1275 | 34 | 105 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT1G75080.2 | 4e-77 | BES1 family protein | ||||



