 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | C.cajan_10973 | ||||||||
| Common Name | KK1_011297 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; fabids; Fabales; Fabaceae; Papilionoideae; Phaseoleae; Cajanus
|
||||||||
| Family | MYB_related | ||||||||
| Protein Properties | Length: 766aa MW: 84241.8 Da PI: 6.4347 | ||||||||
| Description | MYB_related family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | Myb_DNA-binding | 49 | 1.4e-15 | 24 | 68 | 1 | 47 |
TSSS-HHHHHHHHHHHHHTTTT-HHHHHHHHTTTS-HHHHHHHHHHH CS
Myb_DNA-binding 1 rgrWTteEdellvdavkqlGggtWktIartmgkgRtlkqcksrwqky 47
r rWT++E+ ++++a k++G W +I +++g ++t+ q++s+ qk+
C.cajan_10973 24 RERWTEDEHNRFLEALKLYGRA-WQRIEEHIG-TKTAVQIRSHAQKF 68
78******************88.*********.************98 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| SuperFamily | SSF46689 | 7.1E-16 | 18 | 73 | IPR009057 | Homeodomain-like |
| PROSITE profile | PS51294 | 19.15 | 19 | 73 | IPR017930 | Myb domain |
| TIGRFAMs | TIGR01557 | 6.4E-17 | 22 | 71 | IPR006447 | Myb domain, plants |
| SMART | SM00717 | 2.3E-12 | 23 | 71 | IPR001005 | SANT/Myb domain |
| Pfam | PF00249 | 8.4E-13 | 24 | 67 | IPR001005 | SANT/Myb domain |
| Gene3D | G3DSA:1.10.10.60 | 6.7E-8 | 24 | 64 | IPR009057 | Homeodomain-like |
| CDD | cd00167 | 5.07E-9 | 26 | 69 | No hit | No description |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0009409 | Biological Process | response to cold | ||||
| GO:0009651 | Biological Process | response to salt stress | ||||
| GO:0009723 | Biological Process | response to ethylene | ||||
| GO:0009733 | Biological Process | response to auxin | ||||
| GO:0009737 | Biological Process | response to abscisic acid | ||||
| GO:0009739 | Biological Process | response to gibberellin | ||||
| GO:0009751 | Biological Process | response to salicylic acid | ||||
| GO:0009753 | Biological Process | response to jasmonic acid | ||||
| GO:0010243 | Biological Process | response to organonitrogen compound | ||||
| GO:0042754 | Biological Process | negative regulation of circadian rhythm | ||||
| GO:0043496 | Biological Process | regulation of protein homodimerization activity | ||||
| GO:0045892 | Biological Process | negative regulation of transcription, DNA-templated | ||||
| GO:0045893 | Biological Process | positive regulation of transcription, DNA-templated | ||||
| GO:0046686 | Biological Process | response to cadmium ion | ||||
| GO:0048574 | Biological Process | long-day photoperiodism, flowering | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0043565 | Molecular Function | sequence-specific DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 766 aa Download sequence Send to blast |
MDAYSSGEEV VIKTRKPYTI TKQRERWTED EHNRFLEALK LYGRAWQRIE EHIGTKTAVQ 60 IRSHAQKFFS KVNPTDPHIK TMTNTCIFGE PNTKLEKEAF VKGVPIGQAL DIDIPPPRPK 120 RKPNNPYPRK TNVGDPTLHN GVKHGKSLVS IASSHSKQAL DLEKEPLPEK HNKDESCSKV 180 FTIIREAPCS SVSSANKNSL SMSVPLRNSC ALREFIPSIK EIITPDITNE SFITDELENQ 240 KLELDDGKQT QKTNGTCKVS KLENSDASKL IQTAKTEVPN CALTIDGMQS NQNYPRHVPV 300 HVVDGSSTQN PSPDMVFQNS IFQPIGGVNG KPNLVTTSAT SNISESQNNT ARSSIHQSFP 360 LCPLFTQHNQ DDYHSFLHIS STFSSLIVST LLQNPAAHAA ASFAATFWPY ANAETSTDSP 420 VCTPGFPSRQ IGSPPSVTAI AAATVAAATA WWAAHGLLPL CAPPHTAFAC PPASATAVPS 480 MNIGESPQKT EQGEIKPQNP PLQDQIIDPE QSEALQAQHS ASKSPAVSSP ESEESGDANL 540 NTSSKATTNH EMNQTISENP DSNKMKGRKL VDRSSCGSNA TSSSEETELL EKDEKEKEEP 600 KTLEENLLAT ETNNRRSRSI SNLTDSWKEV SEEGRLAFQA LFSREVLPQS FSPPYDLINQ 660 DHQIDRNSFK DNKQNTDYKD EDLESKKCSS NCDGVQKNLP LVKDNSEEEG LLSIGLGQGR 720 LKTRRTGFKP YKRCSVEAKE NMIGTACNQG EEKGPKRIRL NEEAST |
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00654 | PBM | Transfer from LOC_Os08g06110 | Download |
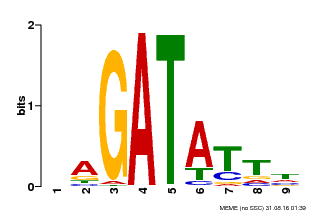
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | C.cajan_10973 |
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | - | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | EU076434 | 0.0 | EU076434.1 Glycine max late elongated hypocotyl and circadian clock associated-1-like protein 2 (LCL2) mRNA, complete cds. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_020211113.1 | 0.0 | protein LHY isoform X1 | ||||
| Refseq | XP_020211114.1 | 0.0 | protein LHY isoform X1 | ||||
| TrEMBL | A0A151TY90 | 0.0 | A0A151TY90_CAJCA; Myb-like protein G | ||||
| STRING | GLYMA19G45030.2 | 0.0 | (Glycine max) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Fabids | OGEF4860 | 25 | 37 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT2G46830.1 | 1e-79 | circadian clock associated 1 | ||||



