 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Bv6_147690_mghj.t1 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; Caryophyllales; Chenopodiaceae; Betoideae; Beta
|
||||||||
| Family | ARR-B | ||||||||
| Protein Properties | Length: 628aa MW: 70778.8 Da PI: 5.3103 | ||||||||
| Description | ARR-B family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | G2-like | 78.9 | 6e-25 | 192 | 247 | 1 | 55 |
G2-like 1 kprlrWtpeLHerFveaveqLG.GsekAtPktilelmkvkgLtlehvkSHLQkYRl 55
k+r++Wt eLH++Fveav+qLG s+k Pk+il+lm+v+ Lt+e+v+SHLQkYRl
Bv6_147690_mghj.t1 192 KARVVWTVELHQKFVEAVNQLGiDSDKIGPKKILDLMNVPRLTRENVASHLQKYRL 247
68*******************7447899***************************8 PP
| |||||||
| 2 | Response_reg | 78.9 | 1.8e-26 | 21 | 129 | 1 | 109 |
EEEESSSHHHHHHHHHHHHHTTCEEEEEESSHHHHHHHHHHHH..ESEEEEESSCTTSEHHHHHHHHHHHTTTSEEEEEESTTTHHHHHHH CS
Response_reg 1 vlivdDeplvrellrqalekegyeevaeaddgeealellkekd..pDlillDiempgmdGlellkeireeepklpiivvtahgeeedalea 89
vl+vdD+p+ +++l+++l+k +y ev++ ++eal+ll++++ +D+++ D++mp+m+G++ll++ e +lp+i+++ ge + + +
Bv6_147690_mghj.t1 21 VLVVDDDPTWLKILEKMLKKCNY-EVTTSGLAREALKLLRDRKdgFDIVISDVNMPDMNGFKLLEHVGLEM-DLPVIMMSVDGETSRVMKG 109
89*********************.***************888889**********************6644.8****************** PP
HHTTESEEEESS--HHHHHH CS
Response_reg 90 lkaGakdflsKpfdpeelvk 109
++ Ga d+l Kp+ ++el +
Bv6_147690_mghj.t1 110 VQHGACDYLLKPIRMKELKN 129
****************9986 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| PIRSF | PIRSF036392 | 1.5E-144 | 1 | 588 | IPR017053 | Response regulator B-type, plant |
| Gene3D | G3DSA:3.40.50.2300 | 4.3E-43 | 18 | 142 | No hit | No description |
| SuperFamily | SSF52172 | 5.41E-35 | 18 | 142 | IPR011006 | CheY-like superfamily |
| SMART | SM00448 | 7.0E-31 | 19 | 131 | IPR001789 | Signal transduction response regulator, receiver domain |
| PROSITE profile | PS50110 | 42.31 | 20 | 135 | IPR001789 | Signal transduction response regulator, receiver domain |
| Pfam | PF00072 | 1.4E-23 | 21 | 129 | IPR001789 | Signal transduction response regulator, receiver domain |
| CDD | cd00156 | 1.33E-29 | 22 | 134 | No hit | No description |
| SuperFamily | SSF46689 | 4.03E-18 | 190 | 251 | IPR009057 | Homeodomain-like |
| Gene3D | G3DSA:1.10.10.60 | 2.6E-28 | 190 | 252 | IPR009057 | Homeodomain-like |
| TIGRFAMs | TIGR01557 | 5.7E-22 | 192 | 247 | IPR006447 | Myb domain, plants |
| Pfam | PF00249 | 6.6E-8 | 194 | 246 | IPR001005 | SANT/Myb domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0000160 | Biological Process | phosphorelay signal transduction system | ||||
| GO:0009735 | Biological Process | response to cytokinin | ||||
| GO:0010082 | Biological Process | regulation of root meristem growth | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 628 aa Download sequence Send to blast |
MIMENGFSSP RSEPFPAGLR VLVVDDDPTW LKILEKMLKK CNYEVTTSGL AREALKLLRD 60 RKDGFDIVIS DVNMPDMNGF KLLEHVGLEM DLPVIMMSVD GETSRVMKGV QHGACDYLLK 120 PIRMKELKNI WQHVVRKRMH EVRDIECIED RSGIDHFHDG HYGGDHTFLK KRKDLDNKHD 180 DKDSNDPSAT KKARVVWTVE LHQKFVEAVN QLGIDSDKIG PKKILDLMNV PRLTRENVAS 240 HLQKYRLYLS RIRKEDKMES SCGETKNPDY SPKESSGSFS LQNSSFGQQN DITSKYVYQE 300 NQLPVQTADS KTFDNDLGSS KYLTEHRIPF STENSDSHTS RNSQIGLNHT YGSIDTSIKH 360 SSFDSTVAGR YPWSAEIPDL KFELEYKPHL QLENNLDQPA APIMQQHIIP DLIEPAPHVS 420 ADLLEPAPQC THTSPGPSIS ETNLGLIDIK PLIAKEGGHD IKGLSPMEST LGSFSVSNDD 480 ACGCAFDMKN QNFSQGVNNI TKEPPLLERS FQTFPVQSGS HLMDAHGLDP SFNVTLQMEK 540 QISCQSSVPG SELDLRNQVL KSESASELLR EDLMFRLLQY GDCTSKDFGL GFSLPEHESV 600 LPLYMPKLDY ENSFDWGEYP PDQGLLFS |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 1irz_A | 1e-18 | 188 | 252 | 1 | 63 | ARR10-B |
| Search in ModeBase | ||||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00010 | PBM | Transfer from AT1G67710 | Download |
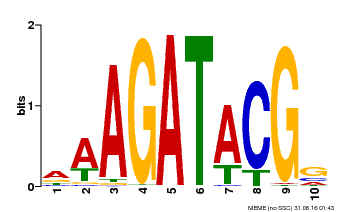
| |||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_010682390.1 | 0.0 | PREDICTED: two-component response regulator ARR11 isoform X1 | ||||
| TrEMBL | A0A0K9QRY8 | 0.0 | A0A0K9QRY8_SPIOL; Two-component response regulator | ||||
| STRING | XP_010682390.1 | 0.0 | (Beta vulgaris) | ||||
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT1G67710.1 | 1e-118 | response regulator 11 | ||||



