 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Brast09G158700.1.p | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; Liliopsida; Petrosaviidae; commelinids; Poales; Poaceae; BOP clade; Pooideae; Brachypodieae; Brachypodium
|
||||||||
| Family | Trihelix | ||||||||
| Protein Properties | Length: 754aa MW: 81036 Da PI: 6.2901 | ||||||||
| Description | Trihelix family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | trihelix | 98.2 | 7e-31 | 87 | 171 | 1 | 87 |
trihelix 1 rWtkqevlaLiearremeerlrrgklkkplWeevskkmrergferspkqCkekwenlnkrykkikegekkrtsessstcpyfdqlea 87
rW+++e+laLi++r+em+ ++r++ lk+plWeevs+k++e g++r++k+Ckek+en++k+yk++keg+ +r++++s +++f++lea
Brast09G158700.1.p 87 RWPREETLALIRIRSEMDATFRDATLKGPLWEEVSRKLAELGYKRNAKKCKEKFENVHKYYKRTKEGRTGRQDGKS--YRFFSELEA 171
8*********************************************************************866665..******985 PP
| |||||||
| 2 | trihelix | 112.4 | 2.6e-35 | 493 | 578 | 1 | 87 |
trihelix 1 rWtkqevlaLiearremeerlrrgklkkplWeevskkmrergferspkqCkekwenlnkrykkikegekkrtsessstcpyfdqlea 87
rW+k ev+aLi++r +m++r++++ k+plWee+s+ mr+ g++r+pk+Ckekwen+nk++kk+ke++k+r +e+s+tcpyf+qlea
Brast09G158700.1.p 493 RWPKTEVHALIQLRMDMDNRYQENGPKGPLWEEISSGMRRLGYNRNPKRCKEKWENINKYFKKVKESNKRR-PEDSKTCPYFHQLEA 578
8********************************************************************97.99***********85 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| SMART | SM00717 | 0.0012 | 84 | 146 | IPR001005 | SANT/Myb domain |
| Pfam | PF13837 | 6.4E-22 | 86 | 172 | No hit | No description |
| CDD | cd12203 | 9.16E-31 | 86 | 151 | No hit | No description |
| PROSITE profile | PS50090 | 6.98 | 86 | 144 | IPR017877 | Myb-like domain |
| SMART | SM00717 | 0.002 | 490 | 552 | IPR001005 | SANT/Myb domain |
| PROSITE profile | PS50090 | 7.236 | 492 | 550 | IPR017877 | Myb-like domain |
| Gene3D | G3DSA:1.10.10.60 | 5.7E-4 | 492 | 549 | IPR009057 | Homeodomain-like |
| Pfam | PF13837 | 4.6E-23 | 492 | 579 | No hit | No description |
| CDD | cd12203 | 7.16E-31 | 492 | 557 | No hit | No description |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 754 aa Download sequence Send to blast |
MQQHQGGGSQ YGAAPPQDMG PFSTPPAAPG PVPLSSRPPP PTQQPTYEEL AAASGAGAGG 60 GFPDDDMLGD SGGNSGGGGG LGASGNRWPR EETLALIRIR SEMDATFRDA TLKGPLWEEV 120 SRKLAELGYK RNAKKCKEKF ENVHKYYKRT KEGRTGRQDG KSYRFFSELE ALHATTAQHH 180 QEQPPPVSGA SGPAPPQLHA FAAPVSAPLP MSTMPPPPGP IQPAPISSAA PAAHVELQQP 240 PPVGLQGLSF PSMSESESDD ESEDDEMTAE TGGGSGSPDG LGKRKHGGGG SSKKMMAFFE 300 GLMKQVIQRQ EEMQRRFLET MEKREAERTA REEAWRKQEV ARLNREQEIL AHERAAAASR 360 DASIIAFLQR VGGGQAVQAP APVVIPMPAP MQVQTTPAAR QPPRQQLPPP PPPPPPQPIP 420 AAPLQQQPPP PQPQHKETAT HQEAVTTPRS APAPAPISGG TSLALVPVGE QQVETHGLGG 480 GGGDHGGAAS SSRWPKTEVH ALIQLRMDMD NRYQENGPKG PLWEEISSGM RRLGYNRNPK 540 RCKEKWENIN KYFKKVKESN KRRPEDSKTC PYFHQLEAIY RKKHNGAGAV SAANAVVASV 600 CAVTTADQQQ TLNRHEIEGK KINDNDKRNN GGVGEGATQT QVPTSNGETT PTTTPPVDFS 660 EKKPEDTVRE LSEQQQREFT TDETDSDDMG DDYTDDGEDG EDDGKMQYRI QFQTPNPVRN 720 PVGTNNAPPP ATTPTNAVPP TSTPSSSFLA MVQ* |
| Nucleic Localization Signal ? help Back to Top | |||
|---|---|---|---|
| No. | Start | End | Sequence |
| 1 | 129 | 137 | KRNAKKCKE |
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Probable transcription factor that binds specific DNA sequence. {ECO:0000250}. | |||||
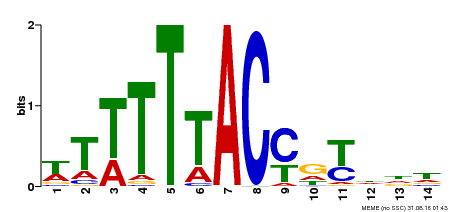
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00244 | DAP | Transfer from AT1G76890 | Download |
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | Brast09G158700.1.p |
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | AK358911 | 1e-138 | AK358911.1 Hordeum vulgare subsp. vulgare mRNA for predicted protein, complete cds, clone: NIASHv1085J16. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_010240190.1 | 0.0 | trihelix transcription factor GTL1 isoform X1 | ||||
| Swissprot | Q39117 | 4e-72 | TGT2_ARATH; Trihelix transcription factor GT-2 | ||||
| TrEMBL | I1J064 | 0.0 | I1J064_BRADI; Uncharacterized protein | ||||
| STRING | BRADI5G17150.1 | 0.0 | (Brachypodium distachyon) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Monocots | OGMP606 | 38 | 175 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT1G76880.1 | 2e-50 | Trihelix family protein | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | Brast09G158700.1.p |



