 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Bostr.0597s0142.1.p | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; malvids; Brassicales; Brassicaceae; Boechereae; Boechera
|
||||||||
| Family | FAR1 | ||||||||
| Protein Properties | Length: 791aa MW: 91338.9 Da PI: 7.3918 | ||||||||
| Description | FAR1 family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | FAR1 | 89.3 | 5.4e-28 | 66 | 154 | 2 | 91 |
FAR1 2 fYneYAkevGFsvrkskskkskrngeitkrtfvCskegkreeekkktekerrtraetrtgCkaklkvkkekdgkwevtkleleHnHelap 91
fY+eYAk++GF++++++s++sk+++e+++++f Cs++g + e++++ + ++r++++++t+Cka+++vk++ dgkw ++++ ++HnHel p
Bostr.0597s0142.1.p 66 FYQEYAKSMGFTTSIKNSRRSKKTKEFIDAKFACSRYGVTPETESS-SCSSRRSSVKKTDCKASMHVKRRPDGKWIIHEFIKDHNHELLP 154
9***************************************999998.88999999********************************975 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| Pfam | PF03101 | 8.0E-26 | 66 | 154 | IPR004330 | FAR1 DNA binding domain |
| Pfam | PF10551 | 6.3E-26 | 275 | 368 | IPR018289 | MULE transposase domain |
| PROSITE profile | PS50966 | 8.939 | 555 | 591 | IPR007527 | Zinc finger, SWIM-type |
| Pfam | PF04434 | 9.0E-7 | 557 | 590 | IPR007527 | Zinc finger, SWIM-type |
| SMART | SM00575 | 3.1E-7 | 566 | 593 | IPR006564 | Zinc finger, PMZ-type |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0010018 | Biological Process | far-red light signaling pathway | ||||
| GO:0042753 | Biological Process | positive regulation of circadian rhythm | ||||
| GO:0045893 | Biological Process | positive regulation of transcription, DNA-templated | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| GO:0008270 | Molecular Function | zinc ion binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 791 aa Download sequence Send to blast |
MELQENQVSD AGDECMVDIM VESNSNRDVG IVADLKTGGD LGFSGVLDLE PRNGIDFESN 60 DAAYLFYQEY AKSMGFTTSI KNSRRSKKTK EFIDAKFACS RYGVTPETES SSCSSRRSSV 120 KKTDCKASMH VKRRPDGKWI IHEFIKDHNH ELLPALAYHF RIQRNVKLAE KNNIDILHAV 180 SERTRKMYVE MSRQSGGYKN IGPLLQTDVN NQVDKGRYLA LEEGDSQVLL EYFKRIKKEN 240 SKFFYAIDLN KEQRLRNLFW ADAKSRDDYL SFNDVVSFDT TYVKFNDKLP LALFIGVNHH 300 SQPMLVGCAL VADESKETFV WLIKTWLRAM GGRAPKVILT DQDKFLMAAV SELLPNTRHC 360 FALWHVLEKI PEYFSHVMKR HENFLQKFNK CIFRSWTDDE FDMRWWKMVS QFGLENDEWL 420 LWLHEHRQKW VPTFMSDVFL AGMSTSQRSE SVNSFFDKYV HKKITLKEFL RLYGAILQNR 480 YEEESVADFD TCHKQPALKS PSPWEKQMAT TYTHTIFKKF QVEVLGVVAC HPRKEKEDEN 540 MATFRVQDCE KDNDFLVTWS KTKSELCCFC RMFEYKGFLC RHALMILQMC GFASIPPRYI 600 LKRWTKDAKS GVLAGEGADQ IQTRVQRYND LCSRAIELSE EGCVSEENYN IVLRTLVESL 660 KNCVDLNNAR NNITESSSQL NNGAHEEENQ VMAGVKATKK KTVVRKRKGQ QEADQMLESQ 720 QSLQPMENIS SEGMSLNGYY GPQQNVQGLL NLMEPPHEGY YVNQRTIQGL GQLNSIAPVQ 780 DSFFANQQAM P |
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
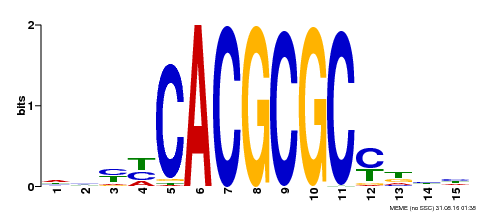
| UniProt | Transcription activator that recognizes and binds to the DNA consensus sequence 5'-CACGCGC-3'. Activates the expression of FHY1 and FHL involved in light responses. Positive regulator of chlorophyll biosynthesis via the activation of HEMB1 gene expression. {ECO:0000269|PubMed:11889039, ECO:0000269|PubMed:12753585, ECO:0000269|PubMed:18033885, ECO:0000269|PubMed:22634759}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00434 | DAP | Transfer from AT4G15090 | Download |
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | Bostr.0597s0142.1.p |
| Regulation -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | INDUCTION: Down-regulated after exposure to far-red light. Subject to a negative feedback regulation by PHYA signaling. {ECO:0000269|PubMed:18033885}. | |||||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | AF159587 | 0.0 | AF159587.1 Arabidopsis thaliana far-red impaired response protein (FAR1) mRNA, complete cds. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_010435013.1 | 0.0 | PREDICTED: protein FAR-RED IMPAIRED RESPONSE 1 | ||||
| Refseq | XP_019086857.1 | 0.0 | PREDICTED: protein FAR-RED IMPAIRED RESPONSE 1 | ||||
| Refseq | XP_020874483.1 | 0.0 | protein FAR-RED IMPAIRED RESPONSE 1 isoform X1 | ||||
| Refseq | XP_020874484.1 | 0.0 | protein FAR-RED IMPAIRED RESPONSE 1 isoform X2 | ||||
| Swissprot | Q9SWG3 | 0.0 | FAR1_ARATH; Protein FAR-RED IMPAIRED RESPONSE 1 | ||||
| TrEMBL | A0A178UY59 | 0.0 | A0A178UY59_ARATH; FAR1 | ||||
| TrEMBL | A0A178UYQ0 | 0.0 | A0A178UYQ0_ARATH; FAR1 | ||||
| STRING | Bostr.0597s0142.1.p | 0.0 | (Boechera stricta) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Malvids | OGEM118 | 26 | 253 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT4G15090.1 | 0.0 | FAR1 family protein | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | Bostr.0597s0142.1.p |



