 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Bathy01g03260 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Chlorophyta; prasinophytes; Mamiellophyceae; Mamiellales; Bathycoccaceae; Bathycoccus
|
||||||||
| Family | ERF | ||||||||
| Protein Properties | Length: 603aa MW: 61925.7 Da PI: 10.2948 | ||||||||
| Description | ERF family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | AP2 | 56.3 | 7.9e-18 | 374 | 424 | 1 | 55 |
AP2 1 sgykGVrwdkkrgrWvAeIrdpsengkr.krfslgkfgtaeeAakaaiaarkkleg 55
s y+GVr+++ +g+++AeIrdp+ r r +lg+f++ae+Aa a++aa+++++g
Bathy01g03260 374 SIYRGVRQRP-WGKFAAEIRDPT----RgSRLWLGTFDSAEDAALAYDAAARAIRG 424
57********.**********84....35***********************9997 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| SuperFamily | SSF54171 | 3.33E-21 | 374 | 432 | IPR016177 | DNA-binding domain |
| CDD | cd00018 | 3.91E-17 | 374 | 432 | No hit | No description |
| Pfam | PF00847 | 1.2E-11 | 374 | 424 | IPR001471 | AP2/ERF domain |
| Gene3D | G3DSA:3.30.730.10 | 5.1E-30 | 375 | 432 | IPR001471 | AP2/ERF domain |
| SMART | SM00380 | 5.5E-34 | 375 | 438 | IPR001471 | AP2/ERF domain |
| PROSITE profile | PS51032 | 22.431 | 375 | 432 | IPR001471 | AP2/ERF domain |
| PRINTS | PR00367 | 9.8E-10 | 376 | 387 | IPR001471 | AP2/ERF domain |
| PRINTS | PR00367 | 9.8E-10 | 398 | 414 | IPR001471 | AP2/ERF domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0008219 | Biological Process | cell death | ||||
| GO:0009414 | Biological Process | response to water deprivation | ||||
| GO:0009723 | Biological Process | response to ethylene | ||||
| GO:0009735 | Biological Process | response to cytokinin | ||||
| GO:0009737 | Biological Process | response to abscisic acid | ||||
| GO:0009753 | Biological Process | response to jasmonic acid | ||||
| GO:0010200 | Biological Process | response to chitin | ||||
| GO:0010286 | Biological Process | heat acclimation | ||||
| GO:0045892 | Biological Process | negative regulation of transcription, DNA-templated | ||||
| GO:0045893 | Biological Process | positive regulation of transcription, DNA-templated | ||||
| GO:0051707 | Biological Process | response to other organism | ||||
| GO:0005730 | Cellular Component | nucleolus | ||||
| GO:0005737 | Cellular Component | cytoplasm | ||||
| GO:0005886 | Cellular Component | plasma membrane | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| GO:0043565 | Molecular Function | sequence-specific DNA binding | ||||
| GO:0044212 | Molecular Function | transcription regulatory region DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 603 aa Download sequence Send to blast |
MGFFNPAATI GFPNMFGTTN TSAGGYQPGG IDPATVAAFA SGFYAHSAPA GKSTGASGRG 60 FLQATTSTSS GNAQQSQQQQ QQQQQQMQQQ MQVLGTSPTF YGSSPTAAPM PSRATIGGGT 120 AAQQNAAAQA AAQAAANFMN NVAAMNNAVA SHQKETLALL QQQQQQNFHN QARRGSSEDS 180 GSFGATAAAN AAAMSNSPGF SLQTPQMPAW TKGLSSSVGN SAGTQVTLGS SPNSGFSTFS 240 PVNHSQRGSA SFNYGHVGAG NATSNQLDKQ QQQQPSVGRR NSTSGRHAFE GRNSSLSASP 300 SSSPPVRGGG VQKSTKNDPS VLKRNNLSNN AAFVEGLASA AAASKAVANA DGGSNDGSKE 360 ATAGKTASGK AQKSIYRGVR QRPWGKFAAE IRDPTRGSRL WLGTFDSAED AALAYDAAAR 420 AIRGDAAVTN FAKDVVIPEH VRAQLPPLPV RDENGNIIET HGHTSGNAPG NAKKGGNSKK 480 QQITPGGLAP EFANSTPTVQ KESKKKNKGG LPSEDASMLM MLSGAALLDD NDSPNTTNDG 540 SKSTNTTEDA EALQDMDMDD AEAINTSTNA PKLNLKGSKT KGKGKSTKNV ADAMTSAMNH 600 KY* |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 2gcc_A | 1e-21 | 376 | 431 | 6 | 62 | ATERF1 |
| 3gcc_A | 1e-21 | 376 | 431 | 6 | 62 | ATERF1 |
| Search in ModeBase | ||||||
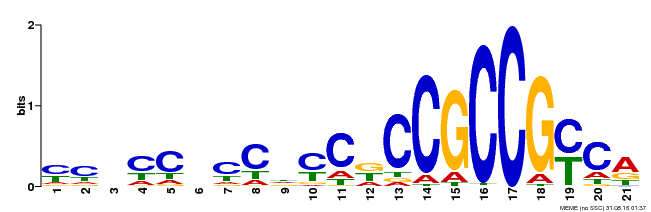
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00366 | DAP | Transfer from AT3G20310 | Download |
| |||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | FO082278 | 0.0 | FO082278.1 Bathycoccus prasinos genomic : Chromosome_1. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_007515094.1 | 0.0 | unknown | ||||
| TrEMBL | K8EXL6 | 0.0 | K8EXL6_9CHLO; Uncharacterized protein | ||||
| STRING | XP_007515094.1 | 0.0 | (Bathycoccus prasinos) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Chlorophytae | OGCP132 | 15 | 40 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT2G33710.1 | 9e-14 | ERF family protein | ||||
| Link Out ? help Back to Top | |
|---|---|
| Entrez Gene | 19018061 |



