 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Araip.4D681 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; fabids; Fabales; Fabaceae; Papilionoideae; Dalbergieae; Arachis
|
||||||||
| Family | MYB_related | ||||||||
| Protein Properties | Length: 401aa MW: 43813.7 Da PI: 8.7622 | ||||||||
| Description | MYB_related family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | Myb_DNA-binding | 39.1 | 1.7e-12 | 189 | 233 | 3 | 47 |
SS-HHHHHHHHHHHHHTTTT-HHHHHHHHTTTS-HHHHHHHHHHH CS
Myb_DNA-binding 3 rWTteEdellvdavkqlGggtWktIartmgkgRtlkqcksrwqky 47
WT+eE++l++ + ++ +g+W+ I+r k+Rt+ q+ s+ qky
Araip.4D681 189 QWTEEEHKLFLVGLQKVRKGDWRGISRNYVKTRTPTQVASHAQKY 233
6*******************************************9 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| PROSITE profile | PS51294 | 16.426 | 182 | 238 | IPR017930 | Myb domain |
| SuperFamily | SSF46689 | 9.36E-16 | 184 | 238 | IPR009057 | Homeodomain-like |
| TIGRFAMs | TIGR01557 | 7.5E-15 | 185 | 236 | IPR006447 | Myb domain, plants |
| SMART | SM00717 | 1.6E-7 | 186 | 236 | IPR001005 | SANT/Myb domain |
| Gene3D | G3DSA:1.10.10.60 | 7.7E-9 | 188 | 233 | IPR009057 | Homeodomain-like |
| Pfam | PF00249 | 5.3E-10 | 189 | 233 | IPR001005 | SANT/Myb domain |
| CDD | cd00167 | 1.60E-7 | 190 | 234 | No hit | No description |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0009651 | Biological Process | response to salt stress | ||||
| GO:0009723 | Biological Process | response to ethylene | ||||
| GO:0009733 | Biological Process | response to auxin | ||||
| GO:0009737 | Biological Process | response to abscisic acid | ||||
| GO:0009739 | Biological Process | response to gibberellin | ||||
| GO:0009751 | Biological Process | response to salicylic acid | ||||
| GO:0009753 | Biological Process | response to jasmonic acid | ||||
| GO:0046686 | Biological Process | response to cadmium ion | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 401 aa Download sequence Send to blast |
MLFGVRVVVD SMRKSVSMNN LSQYVHPQDE SNNNKDAAFS ATGYASADDA APHNSGKNRE 60 RERKRGVPWT EEEHKLFLYG SSDSASSPTQ LARSKKMCST FAAAADSSIS GDGDSPTAVK 120 IMLFGIRVVV DSMRKSVSMN NLSQYVHPQD DSNNNKDAAF STTGYASTDD AAPHNFGKNR 180 ERERKRGVQW TEEEHKLFLV GLQKVRKGDW RGISRNYVKT RTPTQVASHA QKYFLRRSNL 240 NRRRRRSSLF DITTDSVSSI PMEEEQIQNG DSVPHSQPLC PAAPENSNTN GFPMVPVYPL 300 GVGPGVISSV EAGNPMDELT LGQANMELNA QTKLLHPIPV ASNPKTCTVS DVASGSNSPL 360 DLPTLSLGLS FTSDQRQTPS RHSAFHSIPS LNNGDSVISV A |
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Binds selectively to the DNA sequence 5'-[GA]GATAA-3' and may act as a transcription factor involved in the regulation of drought-responsive genes. Enhances stomatal closure in response to abscisic acid (ABA). Confers drought and salt tolerance. {ECO:0000269|PubMed:21030505}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00631 | PBM | Transfer from PK23829.1 | Download |
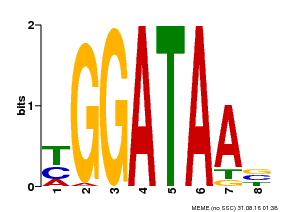
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | Araip.4D681 |
| Regulation -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | INDUCTION: Up-regulated by drought, high salinity, and ABA. {ECO:0000269|PubMed:21030505}. | |||||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | - | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | KF208668 | 0.0 | KF208668.1 Arachis hypogaea putative MYB-related protein 14 (MYB14) gene, complete cds. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_016169474.1 | 0.0 | LOW QUALITY PROTEIN: transcription factor MYB1R1-like | ||||
| Swissprot | Q2V9B0 | 1e-66 | MY1R1_SOLTU; Transcription factor MYB1R1 | ||||
| TrEMBL | A0A444XY95 | 0.0 | A0A444XY95_ARAHY; Uncharacterized protein | ||||
| STRING | XP_007155372.1 | 1e-126 | (Phaseolus vulgaris) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Fabids | OGEF4248 | 31 | 54 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT1G74840.2 | 1e-48 | MYB_related family protein | ||||
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



