 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Araha.57242s0001.1.p | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; malvids; Brassicales; Brassicaceae; Camelineae; Arabidopsis
|
||||||||
| Family | Trihelix | ||||||||
| Protein Properties | Length: 360aa MW: 41209.4 Da PI: 8.454 | ||||||||
| Description | Trihelix family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | trihelix | 86.1 | 4.2e-27 | 264 | 345 | 1 | 85 |
trihelix 1 rWtkqevlaLiearremeerlrrgklkkplWeevskkmrergferspkqCkekwenlnkrykkikegekkrtsessstcpyfdql 85
rW+++ev+aLi r+++ee+ + + k++ W+e+s++m+erg+ers+k+Ckekwen+nk+y++++eg +k+ +e+s+t +yf++l
Araha.57242s0001.1.p 264 RWPQEEVQALISTRSDVEEKTGIN--KGAIWDEISARMKERGYERSAKKCKEKWENMNKYYRRVTEGGGKQ-PEHSKTRSYFEKL 345
8******************99988..99*****************************************95.9**********98 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| SuperFamily | SSF46689 | 3.03E-5 | 262 | 347 | IPR009057 | Homeodomain-like |
| Pfam | PF13837 | 2.9E-20 | 263 | 345 | No hit | No description |
| Gene3D | G3DSA:1.10.10.60 | 6.2E-5 | 263 | 319 | IPR009057 | Homeodomain-like |
| CDD | cd12203 | 9.77E-23 | 263 | 326 | No hit | No description |
| PROSITE profile | PS50090 | 7.503 | 264 | 319 | IPR017877 | Myb-like domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 360 aa Download sequence Send to blast |
SRQSGSHGND DALATLADLA SPPQKLKPIR CGVKLPSEDR HPLDILAGTL DRLPEMGFGC 60 FEAPLGSKIA DVEQSGQLTR GFSKDDDDSL PLQMEFQARN RISWDGLSLS SSVDSDSDSS 120 SDVRKTVTGK RKRETRVKLE HFLEKLVGSM MKRQEKMHNQ LIKVMEKMEG ERIRREEAWR 180 QQEIERMIQN EEARKQELAR SLSLISFIRS VIGDEIEIPK HFEFPQPLQQ ILPEQCKDEK 240 CEPAQTEKEM KFRYSSGSGS SGRRWPQEEV QALISTRSDV EEKTGINKGA IWDEISARMK 300 ERGYERSAKK CKEKWENMNK YYRRVTEGGG KQPEHSKTRS YFEKLGNFYK TNSSGEREK* |
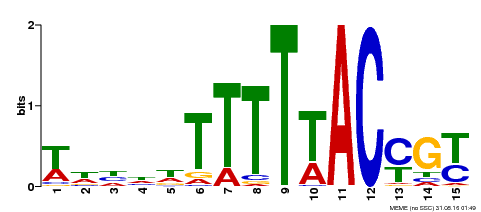
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00552 | DAP | Transfer from AT5G47660 | Download |
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | Araha.57242s0001.1.p |
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | AB025628 | 0.0 | AB025628.1 Arabidopsis thaliana genomic DNA, chromosome 5, P1 clone:MNJ7. | |||
| GenBank | CP002688 | 0.0 | CP002688.1 Arabidopsis thaliana chromosome 5 sequence. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_002865109.1 | 0.0 | trihelix transcription factor GTL1 | ||||
| TrEMBL | D7MP23 | 0.0 | D7MP23_ARALL; Predicted protein | ||||
| STRING | Al_scaffold_0008_21 | 0.0 | (Arabidopsis lyrata) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Malvids | OGEM9411 | 27 | 36 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT5G47660.1 | 0.0 | Trihelix family protein | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | Araha.57242s0001.1.p |



