 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Araha.11372s0005.1.p | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; malvids; Brassicales; Brassicaceae; Camelineae; Arabidopsis
|
||||||||
| Family | SBP | ||||||||
| Protein Properties | Length: 393aa MW: 44004.1 Da PI: 8.4368 | ||||||||
| Description | SBP family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | SBP | 124.2 | 5.7e-39 | 174 | 250 | 1 | 77 |
--SSTT-----TT--HHHHHTT--HHHHT-S-EEETTEEEEE-TTTSSEEETTT--SS--S-STTTT-------S-- CS
SBP 1 lCqvegCeadlseakeyhrrhkvCevhskapvvlvsgleqrfCqqCsrfhelsefDeekrsCrrrLakhnerrrkkq 77
+Cq+++Ce dls+ak yhr+h+vCe+hsk+p+v vsgl +rfCqqCsrfh++sefD++krsCr+rL++hn+rrrk+q
Araha.11372s0005.1.p 174 RCQIDNCELDLSSAKGYHRKHRVCEKHSKCPKVSVSGLVRRFCQQCSRFHDVSEFDDKKRSCRKRLSHHNARRRKPQ 250
6**************************************************************************97 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| Gene3D | G3DSA:4.10.1100.10 | 1.3E-30 | 168 | 236 | IPR004333 | Transcription factor, SBP-box |
| PROSITE profile | PS51141 | 31.317 | 172 | 249 | IPR004333 | Transcription factor, SBP-box |
| SuperFamily | SSF103612 | 5.89E-36 | 173 | 251 | IPR004333 | Transcription factor, SBP-box |
| Pfam | PF03110 | 3.4E-29 | 175 | 248 | IPR004333 | Transcription factor, SBP-box |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0048510 | Biological Process | regulation of timing of transition from vegetative to reproductive phase | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 393 aa Download sequence Send to blast |
MDCSMVSSFQ WDWESLIMSN PSKTENEKRQ QSTEWEIEKG DGIESIVPHF SGLERFSSGS 60 ATSFWHTAVS KSSQSTSIHS SSPEVKRCKL ASESSPGDSC SNVNFVQVKT STAPEVSVAS 120 AESDLCLKLG KRTYSEEYWG RNNNEISAVS MKLLTPSVVT RKNSKCGQSM LVPRCQIDNC 180 ELDLSSAKGY HRKHRVCEKH SKCPKVSVSG LVRRFCQQCS RFHDVSEFDD KKRSCRKRLS 240 HHNARRRKPQ GVFSVNPERL YDRRQHTNML WNELSLNTRS EELCEWGTTY DTKPTQTEKG 300 FTLSFQRGNG SEDQLVASNS RMFSTVQTSG GFPAGKSKFQ LHGEAVGEYS GVLHESQDFH 360 RALSLLSTSS DPLAQPHAQP FSPLCSYDVV PK* |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 1ul4_A | 6e-27 | 175 | 248 | 11 | 84 | squamosa promoter binding protein-like 4 |
| Search in ModeBase | ||||||
| Nucleic Localization Signal ? help Back to Top | |||
|---|---|---|---|
| No. | Start | End | Sequence |
| 1 | 230 | 247 | KKRSCRKRLSHHNARRRK |
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
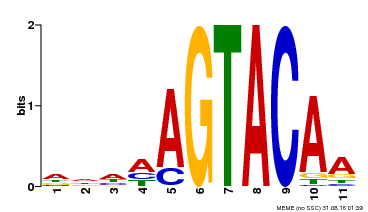
| UniProt | Trans-acting factor that binds specifically to the consensus nucleotide sequence 5'-TNCGTACAA-3'. {ECO:0000250}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00161 | DAP | Transfer from AT1G27360 | Download |
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | Araha.11372s0005.1.p |
| Regulation -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | INDUCTION: Negatively regulated by microRNAs miR156 and miR157. {ECO:0000305|PubMed:12202040}. | |||||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | AJ011635 | 0.0 | AJ011635.1 Arabidopsis thaliana (ecotype Columbia) mRNA for squamosa promoter binding protein-like 11. | |||
| GenBank | AY070106 | 0.0 | AY070106.1 Arabidopsis thaliana putative squamosa-promoter binding protein 2 (At1g27360) mRNA, complete cds. | |||
| GenBank | AY088185 | 0.0 | AY088185.1 Arabidopsis thaliana clone 42666 mRNA, complete sequence. | |||
| GenBank | AY091286 | 0.0 | AY091286.1 Arabidopsis thaliana putative squamosa-promoter binding protein 2 (At1g27360) mRNA, complete cds. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_002890724.1 | 0.0 | squamosa promoter-binding-like protein 11 | ||||
| Refseq | XP_020867910.1 | 0.0 | squamosa promoter-binding-like protein 11 | ||||
| Refseq | XP_020867912.1 | 0.0 | squamosa promoter-binding-like protein 11 | ||||
| Refseq | XP_020867913.1 | 0.0 | squamosa promoter-binding-like protein 11 | ||||
| Swissprot | Q9FZK0 | 0.0 | SPL11_ARATH; Squamosa promoter-binding-like protein 11 | ||||
| TrEMBL | D7KBP3 | 0.0 | D7KBP3_ARALL; Uncharacterized protein | ||||
| STRING | scaffold_103051.1 | 0.0 | (Arabidopsis lyrata) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Malvids | OGEM5004 | 16 | 53 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT1G27360.4 | 0.0 | squamosa promoter-like 11 | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | Araha.11372s0005.1.p |
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



