 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Aradu.ML9JR | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; fabids; Fabales; Fabaceae; Papilionoideae; Dalbergieae; Arachis
|
||||||||
| Family | MYB | ||||||||
| Protein Properties | Length: 988aa MW: 110048 Da PI: 4.9039 | ||||||||
| Description | MYB family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | Myb_DNA-binding | 51.1 | 3e-16 | 36 | 83 | 1 | 48 |
TSSS-HHHHHHHHHHHHHTTTT-HHHHHHHHTTTS-HHHHHHHHHHHT CS
Myb_DNA-binding 1 rgrWTteEdellvdavkqlGggtWktIartmgkgRtlkqcksrwqkyl 48
+g+WT Ed +lvd+v+++G g+W+++ r g+ R++k+c++rw ++l
Aradu.ML9JR 36 KGPWTSIEDAILVDYVTKHGEGNWNAVQRNSGLARCGKSCRLRWANHL 83
79******************************************9986 PP
| |||||||
| 2 | Myb_DNA-binding | 50.8 | 3.8e-16 | 89 | 132 | 1 | 46 |
TSSS-HHHHHHHHHHHHHTTTT-HHHHHHHHTTTS-HHHHHHHHHH CS
Myb_DNA-binding 1 rgrWTteEdellvdavkqlGggtWktIartmgkgRtlkqcksrwqk 46
+g++T+eE+ +++++++++G++ W++ a++++ gRt++++k++w++
Aradu.ML9JR 89 KGAFTEEEERRIIELHAKMGNK-WARMAAELP-GRTDNEIKNYWNT 132
799*******************.*********.***********96 PP
| |||||||
| 3 | Myb_DNA-binding | 52.7 | 9.8e-17 | 503 | 550 | 1 | 48 |
TSSS-HHHHHHHHHHHHHTTTT-HHHHHHHHTTTS-HHHHHHHHHHHT CS
Myb_DNA-binding 1 rgrWTteEdellvdavkqlGggtWktIartmgkgRtlkqcksrwqkyl 48
+g+WT Ed +lvd+vk+ G g+W+++ r+ g+ R++k+c++rw ++l
Aradu.ML9JR 503 KGPWTSAEDAILVDYVKKNGEGNWNAVQRHSGLARCGKSCRLRWANHL 550
79******************************************9986 PP
| |||||||
| 4 | Myb_DNA-binding | 50.8 | 3.8e-16 | 556 | 599 | 1 | 46 |
TSSS-HHHHHHHHHHHHHTTTT-HHHHHHHHTTTS-HHHHHHHHHH CS
Myb_DNA-binding 1 rgrWTteEdellvdavkqlGggtWktIartmgkgRtlkqcksrwqk 46
+g++T+eE+ +++++++++G++ W++ a++++ gRt++++k++w++
Aradu.ML9JR 556 KGAFTEEEERRIIELHAKMGNK-WARMAAELP-GRTDNEIKNYWNT 599
799*******************.*********.***********96 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| PROSITE profile | PS51294 | 13.888 | 31 | 83 | IPR017930 | Myb domain |
| SuperFamily | SSF46689 | 1.57E-28 | 34 | 130 | IPR009057 | Homeodomain-like |
| SMART | SM00717 | 2.6E-12 | 35 | 85 | IPR001005 | SANT/Myb domain |
| Pfam | PF00249 | 1.0E-12 | 36 | 83 | IPR001005 | SANT/Myb domain |
| Gene3D | G3DSA:1.10.10.60 | 1.9E-22 | 37 | 90 | IPR009057 | Homeodomain-like |
| CDD | cd00167 | 1.57E-9 | 38 | 83 | No hit | No description |
| PROSITE profile | PS51294 | 25.573 | 84 | 138 | IPR017930 | Myb domain |
| SMART | SM00717 | 2.3E-16 | 88 | 136 | IPR001005 | SANT/Myb domain |
| Pfam | PF00249 | 4.4E-15 | 89 | 132 | IPR001005 | SANT/Myb domain |
| Gene3D | G3DSA:1.10.10.60 | 5.0E-25 | 91 | 136 | IPR009057 | Homeodomain-like |
| CDD | cd00167 | 3.67E-6 | 103 | 132 | No hit | No description |
| PROSITE profile | PS51294 | 15.549 | 498 | 550 | IPR017930 | Myb domain |
| SuperFamily | SSF46689 | 6.36E-29 | 501 | 597 | IPR009057 | Homeodomain-like |
| SMART | SM00717 | 5.5E-13 | 502 | 552 | IPR001005 | SANT/Myb domain |
| Pfam | PF00249 | 5.8E-14 | 503 | 550 | IPR001005 | SANT/Myb domain |
| Gene3D | G3DSA:1.10.10.60 | 2.1E-22 | 504 | 557 | IPR009057 | Homeodomain-like |
| CDD | cd00167 | 4.68E-10 | 505 | 550 | No hit | No description |
| PROSITE profile | PS51294 | 25.415 | 551 | 605 | IPR017930 | Myb domain |
| SMART | SM00717 | 5.1E-16 | 555 | 603 | IPR001005 | SANT/Myb domain |
| Pfam | PF00249 | 4.4E-15 | 556 | 599 | IPR001005 | SANT/Myb domain |
| Gene3D | G3DSA:1.10.10.60 | 1.1E-24 | 558 | 603 | IPR009057 | Homeodomain-like |
| CDD | cd00167 | 3.67E-6 | 570 | 599 | No hit | No description |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0009789 | Biological Process | positive regulation of abscisic acid-activated signaling pathway | ||||
| GO:0043068 | Biological Process | positive regulation of programmed cell death | ||||
| GO:0045893 | Biological Process | positive regulation of transcription, DNA-templated | ||||
| GO:0045926 | Biological Process | negative regulation of growth | ||||
| GO:0048235 | Biological Process | pollen sperm cell differentiation | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 988 aa Download sequence Send to blast |
MSSMTNMGDG RKVSKSRQSS SSDEEAKTGG EGHLKKGPWT SIEDAILVDY VTKHGEGNWN 60 AVQRNSGLAR CGKSCRLRWA NHLRPDLKKG AFTEEEERRI IELHAKMGNK WARMAAELPG 120 RTDNEIKNYW NTRIKRMQRS GLPIYPPDIC FRVMHGNQES QNVGRLTDGV NQHDDLSQTD 180 TLNIPELDFK HVKIPLDPSI FDIPENSMFE QSSDSSHGYN MFPIMHPAKR LRESAEIMQY 240 GPVYEEDYMS DEITNQFHGN TCCSSSSSMP LSGGMKLELP SLQYLETQQG IWDTPASPLP 300 SLESVDTLSS GLLEPIMYQP MKHFQVQETP DEAVKCKPEW DESGEPYSPL AQSSASVLTM 360 CSPDGPQSVE NSDHNIKHEL GTEFPTYCSG KDEKNMNQIH VTRPDALLDL AWFENSVVNC 420 DGESFRKDAL NTFFDHPTAA DLDVITSTTA VTLYTPLLAK VYSMSTMTNK CDGRKVSKVR 480 RPSSSVEEDS GESNVGGEGH LKKGPWTSAE DAILVDYVKK NGEGNWNAVQ RHSGLARCGK 540 SCRLRWANHL RPDLRKGAFT EEEERRIIEL HAKMGNKWAR MAAELPGRTD NEIKNYWNTR 600 IKRMQRAGLP IYPPDVCLRV MHGNQESVNV GTLRDEANQH DDLSQTDTLD IPELDFKLVK 660 LCPGLSYGTS ICDIPESSMV EMSSDSSHGY NHMFPVMPPS KRLRESDMQY GYLDSCISDS 720 VPIFEQYGDY TCDKISDHTQ LSSPCDPILN TSDQFHGDDT YGSHAPLNGN TSSSSVPLPG 780 AMKLELPSLQ YFETQHGIWG TPASPLPSLE SVDTMIQSPP AKPSPSDPVS PERSGLLEAI 840 IYQPKQLQVA NNNSLQATPN DCVRKEVKIS TMKCKPECDE QGGPHFPLGQ SAASVATNYT 900 TSISMCLVDA PQSVETIQDH NIKQELGTQY PAYVSGKKEN WNQIDFTRPD ALLDLGWFGS 960 GIDCDDYDGD QFLFKDALSP FLGQDPDG |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 1a5j_A | 1e-30 | 34 | 603 | 5 | 106 | B-MYB |
| Search in ModeBase | ||||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00490 | DAP | Transfer from AT5G06100 | Download |
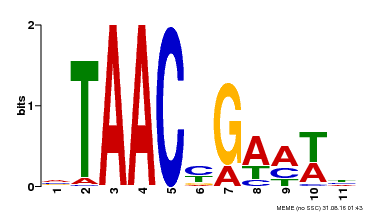
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | Aradu.ML9JR |
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_015958481.1 | 0.0 | transcription factor MYB33 isoform X1 | ||||
| Refseq | XP_015958482.1 | 0.0 | transcription factor MYB33 isoform X1 | ||||
| Refseq | XP_020995322.1 | 0.0 | transcription factor MYB33 isoform X1 | ||||
| TrEMBL | A0A445DKW6 | 0.0 | A0A445DKW6_ARAHY; Uncharacterized protein | ||||
| STRING | GLYMA13G04031.2 | 0.0 | (Glycine max) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Fabids | OGEF2047 | 34 | 85 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT5G06100.3 | 9e-90 | myb domain protein 33 | ||||



